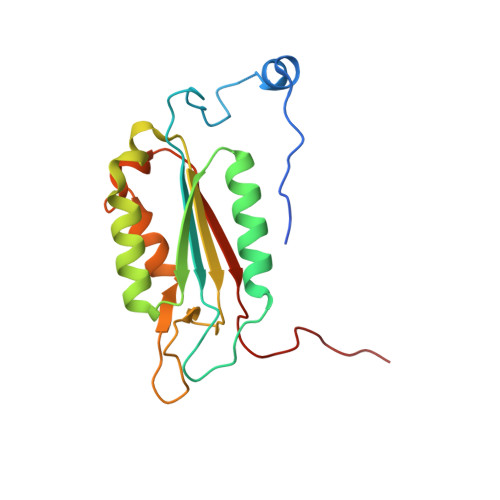
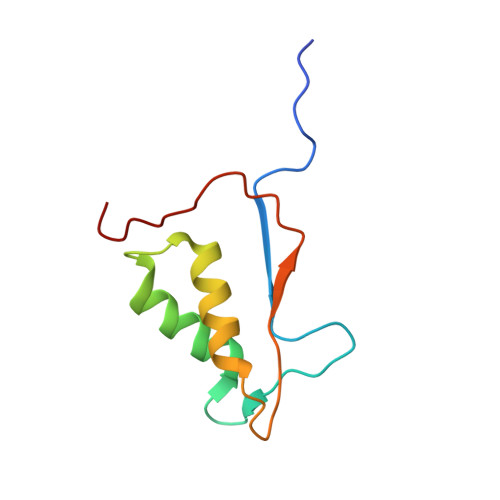
Crystal structure of human Caspase-1 with 2-((2-naphthoyl)-L-valyl)-4-hydroxy-N-((3S)-2-hydroxy-5-oxotetrahydrofuran-3-yl)-2-azabicyclo[2.2.2]octane-3-carboxamide (Compound 1)
Brethon, A., Chantalat, L., Christin, O., Clary, L., Fournier, J.F., Gastreich, M., Harris, C., Pascau, J., Isabet, T., Rodeschin, V., Thoreau, E., Roche, D.To be published.