Structural basis for spumavirus GAG tethering to chromatin.
Lesbats, P., Serrao, E., Maskell, D.P., Pye, V.E., O'Reilly, N., Lindemann, D., Engelman, A.N., Cherepanov, P.(2017) Proc Natl Acad Sci U S A 114: 5509-5514
- PubMed: 28490494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1621159114
- Primary Citation Related Structures:
5MLU - PubMed Abstract:
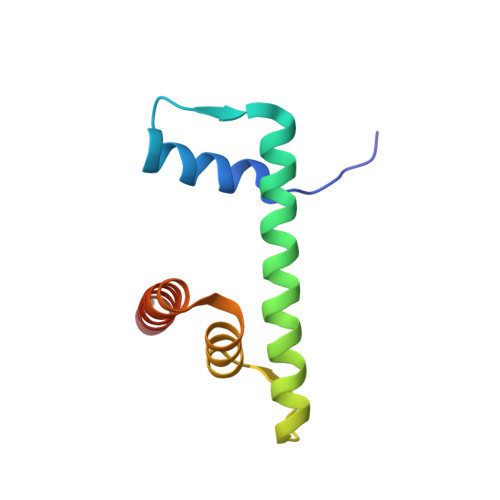
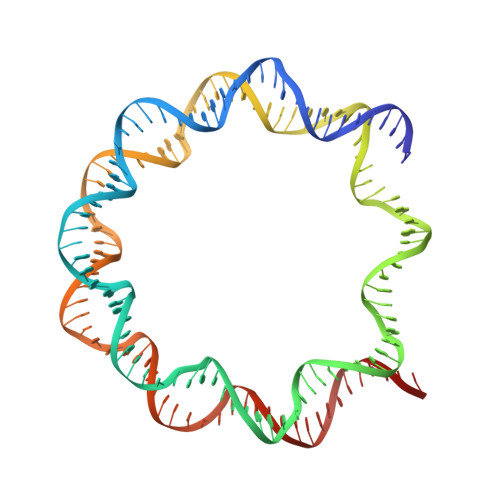
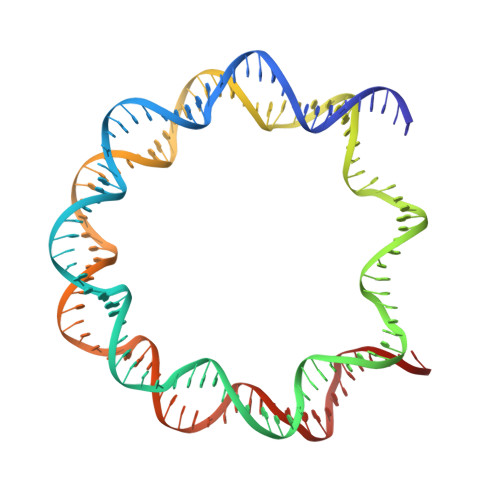
The interactions between a retrovirus and host cell chromatin that underlie integration and provirus expression are poorly understood. The prototype foamy virus (PFV) structural protein GAG associates with chromosomes via a chromatin-binding sequence (CBS) located within its C-terminal region. Here, we show that the PFV CBS is essential and sufficient for a direct interaction with nucleosomes and present a crystal structure of the CBS bound to a mononucleosome. The CBS interacts with the histone octamer, engaging the H2A-H2B acidic patch in a manner similar to other acidic patch-binding proteins such as herpesvirus latency-associated nuclear antigen (LANA). Substitutions of the invariant arginine anchor residue in GAG result in global redistribution of PFV and macaque simian foamy virus (SFV mac ) integration sites toward centromeres, dampening the resulting proviral expression without affecting the overall efficiency of integration. Our findings underscore the importance of retroviral structural proteins for integration site selection and the avoidance of genomic junkyards.
- Chromatin Structure and Mobile DNA, The Francis Crick Institute, London NW1 1AT, United Kingdom.
Organizational Affiliation: