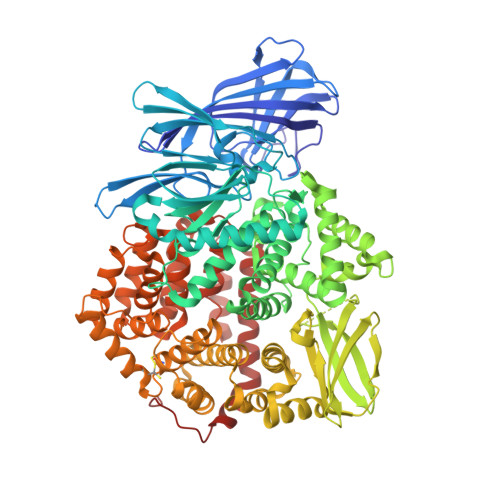
Ligand-Induced Conformational Change of Insulin-Regulated Aminopeptidase: Insights on Catalytic Mechanism and Active Site Plasticity.
Mpakali, A., Saridakis, E., Harlos, K., Zhao, Y., Kokkala, P., Georgiadis, D., Giastas, P., Papakyriakou, A., Stratikos, E.(2017) J Med Chem 60: 2963-2972
- PubMed: 28328206 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01890
- Primary Citation Related Structures:
5MJ6 - PubMed Abstract:
Insulin-regulated aminopeptidase (IRAP) is an enzyme with several important biological functions that is known to process a large variety of different peptidic substrates, although the mechanism behind this wide specificity is not clearly understood. We describe a crystal structure of IRAP in complex with a recently developed bioactive and selective inhibitor at 2.53 Å resolution. In the presence of this inhibitor, the enzyme adopts a novel conformation in which domains II and IV are juxtaposed, forming a hollow structure that excludes external solvent access to the catalytic center. A loop adjacent to the enzyme's GAMEN motif undergoes structural reconfiguration, allowing the accommodation of bulky inhibitor side chains. Atomic interactions between the inhibitor and IRAP that are unique to this conformation can explain the strong selectivity compared to homologous aminopeptidases ERAP1 and ERAP2. This conformation provides insight on IRAP's catalytic cycle and reveals significant active-site plasticity that may underlie its substrate permissiveness.
- National Center for Scientific Research Demokritos, Agia Paraskevi , Athens 15310, Greece.
Organizational Affiliation: