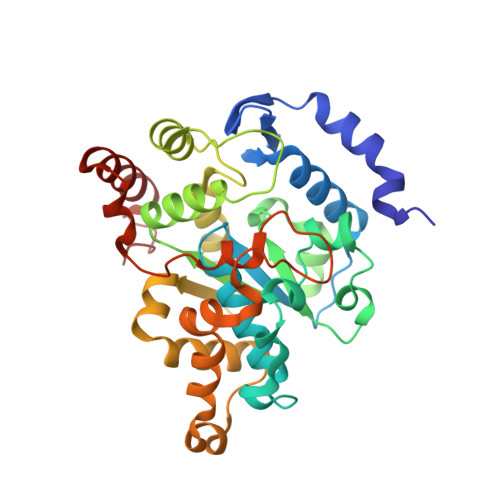
Structural and biochemical studies of sulphotransferase 18 from Arabidopsis thaliana explain its substrate specificity and reaction mechanism.
Hirschmann, F., Krause, F., Baruch, P., Chizhov, I., Mueller, J.W., Manstein, D.J., Papenbrock, J., Fedorov, R.(2017) Sci Rep 7: 4160-4160
- PubMed: 28646214
- DOI: https://doi.org/10.1038/s41598-017-04539-2
- Primary Citation Related Structures:
5MEK, 5MEX - PubMed Abstract:
Sulphotransferases are a diverse group of enzymes catalysing the transfer of a sulfuryl group from 3'-phosphoadenosine 5'-phosphosulphate (PAPS) to a broad range of secondary metabolites. They exist in all kingdoms of life. In Arabidopsis thaliana (L.) Heynh. twenty-two sulphotransferase (SOT) isoforms were identified. Three of those are involved in glucosinolate (Gl) biosynthesis, glycosylated sulphur-containing aldoximes containing chemically different side chains, whose break-down products are involved in stress response against herbivores, pathogens, and abiotic stress. To explain the differences in substrate specificity of desulpho (ds)-Gl SOTs and to understand the reaction mechanism of plant SOTs, we determined the first high-resolution crystal structure of the plant ds-Gl SOT AtSOT18 in complex with 3'-phosphoadenosine 5'-phosphate (PAP) alone and together with the Gl sinigrin. These new structural insights into the determination of substrate specificity were complemented by mutagenesis studies. The structure of AtSOT18 invigorates the similarity between plant and mammalian sulphotransferases, which illustrates the evolutionary conservation of this multifunctional enzyme family. We identified the essential residues for substrate binding and catalysis and demonstrated that the catalytic mechanism is conserved between human and plant enzymes. Our study indicates that the loop-gating mechanism is likely to be a source of the substrate specificity in plants.
- Institute of Botany, Leibniz University Hannover, Herrenhäuserstr. 2, D-30419, Hannover, Germany.
Organizational Affiliation: