Phenylthiourea Binding to Human Tyrosinase-Related Protein 1
Lai, X., Wichers, H.J., Soler-Lopez, M., Dijkstra, B.W.(2020) Int J Mol Sci
Experimental Data Snapshot
(2020) Int J Mol Sci
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 5,6-dihydroxyindole-2-carboxylic acid oxidase | 446 | Homo sapiens | Mutation(s): 3 Gene Names: TYRP1, CAS2, TYRP, TYRRP EC: 1.14.18 | 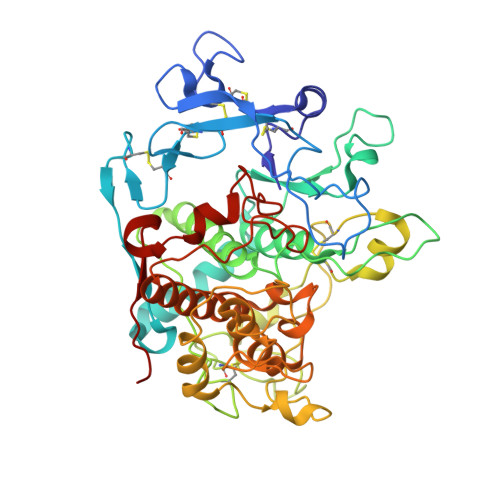 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P17643 GTEx: ENSG00000107165 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P17643 | ||||
Glycosylation | |||||
| Glycosylation Sites: 5 | Go to GlyGen: P17643-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | E, H, P | 3 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G21290RB GlyCosmos: G21290RB GlyGen: G21290RB | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | F, G, J, L, N F, G, J, L, N, O, R, T | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G42666HT GlyCosmos: G42666HT GlyGen: G42666HT | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-6)-2-acetamido-2-deoxy-beta-D-glucopyranose | I | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G87612PG GlyCosmos: G87612PG GlyGen: G87612PG | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | K | 6 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G64181RO GlyCosmos: G64181RO GlyGen: G64181RO | |||||
Entity ID: 6 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-L-fucopyranose-(1-6)-2-acetamido-2-deoxy-beta-D-glucopyranose | M | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G86851RC GlyCosmos: G86851RC GlyGen: G86851RC | |||||
Entity ID: 7 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-3)-alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | Q | 5 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G26032DN GlyCosmos: G26032DN GlyGen: G26032DN | |||||
Entity ID: 8 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | S | 5 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G21381MC GlyCosmos: G21381MC GlyGen: G21381MC | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | CA [auth C], DA [auth C], EA [auth C], U [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| URS Download:Ideal Coordinates CCD File | FA [auth C], IA [auth D], V [auth A], Z [auth B] | N-PHENYLTHIOUREA C7 H8 N2 S FULZLIGZKMKICU-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | AA [auth B] BA [auth B] GA [auth C] HA [auth C] JA [auth D] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 90.081 | α = 90 |
| b = 141.767 | β = 90 |
| c = 191.73 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| PHENIX | refinement |
| SCALA | data scaling |
| PHENIX | phasing |