Design and Synthesis of a Series of l-trans-4-Substituted Prolines as Selective Antagonists for the Ionotropic Glutamate Receptors Including Functional and X-ray Crystallographic Studies of New Subtype Selective Kainic Acid Receptor Subtype 1 (GluK1) Antagonist (2S,4R)-4-(2-Carboxyphenoxy)pyrrolidine-2-carboxylic Acid.
Krogsgaard-Larsen, N., Delgar, C.G., Koch, K., Brown, P.M., Moller, C., Han, L., Huynh, T.H., Hansen, S.W., Nielsen, B., Bowie, D., Pickering, D.S., Kastrup, J.S., Frydenvang, K., Bunch, L.(2017) J Med Chem 60: 441-457
- PubMed: 28005385 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01516
- Primary Citation Related Structures:
5M2V - PubMed Abstract:
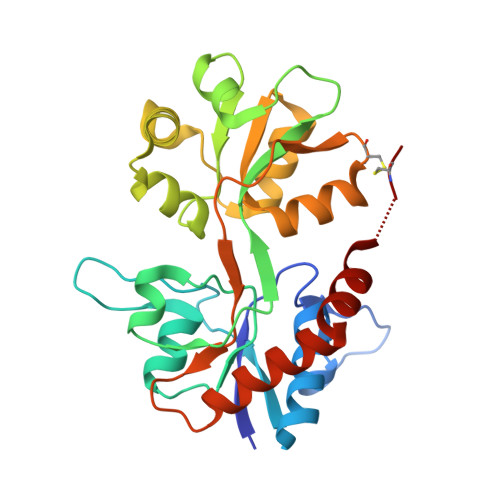
Ionotropic glutamate receptor antagonists are valuable tool compounds for studies of neurological pathways in the central nervous system. On the basis of rational ligand design, a new class of selective antagonists, represented by (2S,4R)-4-(2-carboxyphenoxy)pyrrolidine-2-carboxylic acid (1b), for cloned homomeric kainic acid receptors subtype 1 (GluK1) was attained (K i = 4 μM). In a functional assay, 1b displayed full antagonist activity with IC 50 = 6 ± 2 μM. A crystal structure was obtained of 1b when bound in the ligand binding domain of GluK1. A domain opening of 13-14° was seen compared to the structure with glutamate, consistent with 1b being an antagonist. A structure-activity relationship study showed that the chemical nature of the tethering atom (C, O, or S) linking the pyrrolidine ring and the phenyl ring plays a key role in the receptor selectivity profile and that substituents on the phenyl ring are well accommodated by the GluK1 receptor.
- Bowie Lab, Department of Pharmacology & Therapeutics, Faculty of Medicine, McGill University , Montreal, Quebec H3G 0B1, Canada.
Organizational Affiliation: