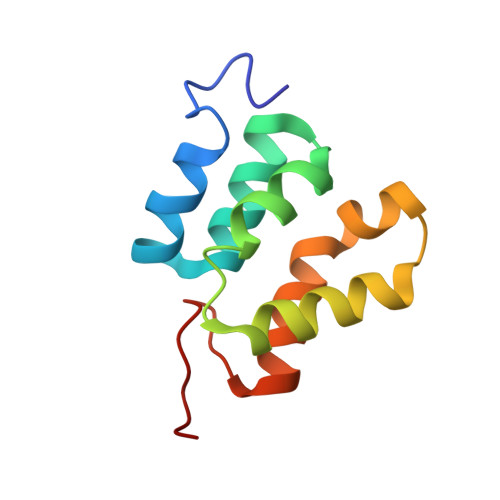
Structure of a Spumaretrovirus Gag Central Domain Reveals an Ancient Retroviral Capsid.
Ball, N.J., Nicastro, G., Dutta, M., Pollard, D.J., Goldstone, D.C., Sanz-Ramos, M., Ramos, A., Mullers, E., Stirnnagel, K., Stanke, N., Lindemann, D., Stoye, J.P., Taylor, W.R., Rosenthal, P.B., Taylor, I.A.(2016) PLoS Pathog 12: e1005981-e1005981
- PubMed: 27829070 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1005981
- Primary Citation Related Structures:
5M1G, 5M1H - PubMed Abstract:
The Spumaretrovirinae, or foamy viruses (FVs) are complex retroviruses that infect many species of monkey and ape. Despite little sequence homology, FV and orthoretroviral Gag proteins perform equivalent functions, including genome packaging, virion assembly, trafficking and membrane targeting. However, there is a paucity of structural information for FVs and it is unclear how disparate FV and orthoretroviral Gag molecules share the same function. To probe the functional overlap of FV and orthoretroviral Gag we have determined the structure of a central region of Gag from the Prototype FV (PFV). The structure comprises two all α-helical domains NtDCEN and CtDCEN that although they have no sequence similarity, we show they share the same core fold as the N- (NtDCA) and C-terminal domains (CtDCA) of archetypal orthoretroviral capsid protein (CA). Moreover, structural comparisons with orthoretroviral CA align PFV NtDCEN and CtDCEN with NtDCA and CtDCA respectively. Further in vitro and functional virological assays reveal that residues making inter-domain NtDCEN-CtDCEN interactions are required for PFV capsid assembly and that intact capsid is required for PFV reverse transcription. These data provide the first information that relates the Gag proteins of Spuma and Orthoretrovirinae and suggests a common ancestor for both lineages containing an ancient CA fold.
- Macromolecular Structure Laboratory, The Francis Crick Institute, Mill Hill Laboratory, London, United Kingdom.
Organizational Affiliation: