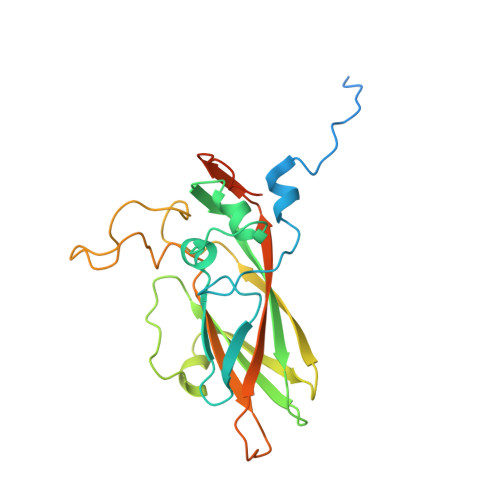
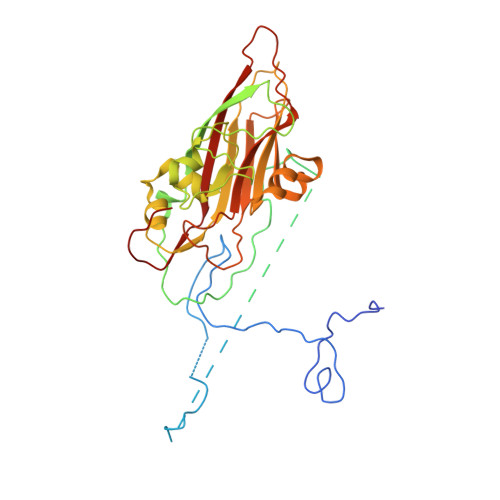
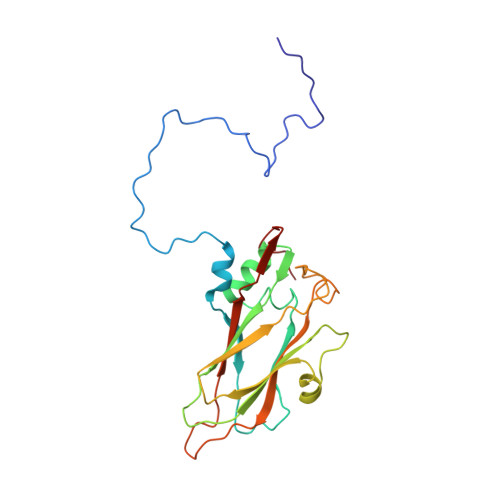
Structure of Aichi Virus 1 and Its Empty Particle: Clues to Kobuvirus Genome Release Mechanism.
Sabin, C., Fuzik, T., Skubnik, K., Palkova, L., Lindberg, A.M., Plevka, P.(2016) J Virol 90: 10800-10810
- PubMed: 27681122 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01601-16
- Primary Citation Related Structures:
5LVC - PubMed Abstract:
Aichi virus 1 (AiV-1) is a human pathogen from the Kobuvirus genus of the Picornaviridae family. Worldwide, 80 to 95% of adults have antibodies against the virus. AiV-1 infections are associated with nausea, gastroenteritis, and fever. Unlike most picornaviruses, kobuvirus capsids are composed of only three types of subunits: VP0, VP1, and VP3. We present here the structure of the AiV-1 virion determined to a resolution of 2.1 Å using X-ray crystallography. The surface loop puff of VP0 and knob of VP3 in AiV-1 are shorter than those in other picornaviruses. Instead, the 42-residue BC loop of VP0 forms the most prominent surface feature of the AiV-1 virion. We determined the structure of AiV-1 empty particle to a resolution of 4.2 Å using cryo-electron microscopy. The empty capsids are expanded relative to the native virus. The N-terminal arms of capsid proteins VP0, which mediate contacts between the pentamers of capsid protein protomers in the native AiV-1 virion, are disordered in the empty capsid. Nevertheless, the empty particles are stable, at least in vitro , and do not contain pores that might serve as channels for genome release. Therefore, extensive and probably reversible local reorganization of AiV-1 capsid is required for its genome release. IMPORTANCE Aichi virus 1 (AiV-1) is a human pathogen that can cause diarrhea, abdominal pain, nausea, vomiting, and fever. AiV-1 is identified in environmental screening studies with higher frequency and greater abundance than other human enteric viruses. Accordingly, 80 to 95% of adults worldwide have suffered from AiV-1 infections. We determined the structure of the AiV-1 virion. Based on the structure, we show that antiviral compounds that were developed against related enteroviruses are unlikely to be effective against AiV-1. The surface of the AiV-1 virion has a unique topology distinct from other related viruses from the Picornaviridae family. We also determined that AiV-1 capsids form compact shells even after genome release. Therefore, AiV-1 genome release requires large localized and probably reversible reorganization of the capsid.
- Structural Virology, Central European Institute of Technology, Masaryk University, Brno, Czech Republic.
Organizational Affiliation: