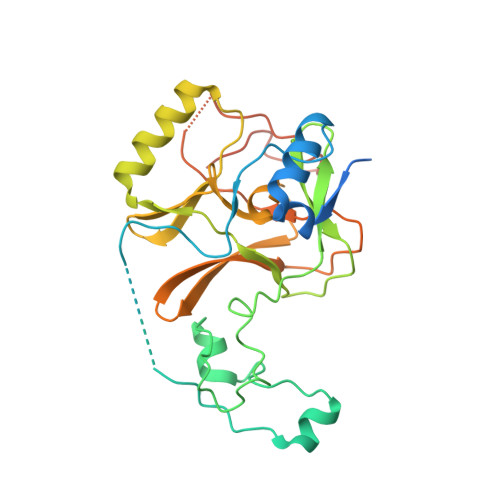
Structure of the Epigenetic Oncogene MMSET and Inhibition by N-Alkyl Sinefungin Derivatives.
Tisi, D., Chiarparin, E., Tamanini, E., Pathuri, P., Coyle, J.E., Hold, A., Holding, F.P., Amin, N., Martin, A.C., Rich, S.J., Berdini, V., Yon, J., Acklam, P., Burke, R., Drouin, L., Harmer, J.E., Jeganathan, F., van Montfort, R.L., Newbatt, Y., Tortorici, M., Westlake, M., Wood, A., Hoelder, S., Heightman, T.D.(2016) ACS Chem Biol 11: 3093-3105
- PubMed: 27571355 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.6b00308
- Primary Citation Related Structures:
5LSS, 5LSU, 5LSX, 5LSY, 5LSZ, 5LT6, 5LT7, 5LT8 - PubMed Abstract:
The members of the NSD subfamily of lysine methyl transferases are compelling oncology targets due to the recent characterization of gain-of-function mutations and translocations in several hematological cancers. To date, these proteins have proven intractable to small molecule inhibition. Here, we present initial efforts to identify inhibitors of MMSET (aka NSD2 or WHSC1) using solution phase and crystal structural methods. On the basis of 2D NMR experiments comparing NSD1 and MMSET structural mobility, we designed an MMSET construct with five point mutations in the N-terminal helix of its SET domain for crystallization experiments and elucidated the structure of the mutant MMSET SET domain at 2.1 Å resolution. Both NSD1 and MMSET crystal systems proved resistant to soaking or cocrystallography with inhibitors. However, use of the close homologue SETD2 as a structural surrogate supported the design and characterization of N-alkyl sinefungin derivatives, which showed low micromolar inhibition against both SETD2 and MMSET.
- Astex Pharmaceuticals , 436 Cambridge Science Park, Milton Road, Cambridge, United Kingdom CB4 0QA.
Organizational Affiliation: