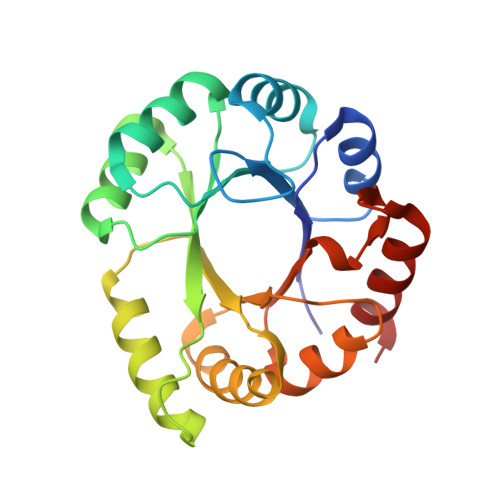
Crystal structure of a phosphoribosyl anthranilate isomerase from the hyperthermophilic archaeon Thermococcus kodakaraensis.
Perveen, S., Rashid, N., Papageorgiou, A.C.(2016) Acta Crystallogr F Struct Biol Commun 72: 804-812
- PubMed: 27827353
- DOI: https://doi.org/10.1107/S2053230X16015223
- Primary Citation Related Structures:
5LHE, 5LHF - PubMed Abstract:
A phosphoribosyl anthranilate isomerase, TkTrpF, from Thermococcus kodakaraensis was expressed in Escherichia coli and purified to homogeneity. TkTrpF was crystallized and its structure was determined by molecular replacement in two different space groups (C2 and P1) using data to 1.85 and 1.75 Å resolution, respectively. TkTrpF belongs to the class of TIM-barrel proteins. Structural comparison with other phosphoribosyl anthranilate isomerases (TrpFs) showed the highest structural similarity to Pyrococcus furiosus TrpF. Similarly to P. furiosus TrpF, TkTrpF is a monomer in solution, in contrast to other thermophilic enzymes, which exist as functional dimers. Although in space group P1 TkTrpF crystallizes with two molecules in the asymmetric unit, the interface is highly improbable in solution. Potential factors for the thermostability of TkTrpF were attributed to an increase in helical structure, an increased number of charged residues and an increase in the number of salt bridges.
- School of Biological Sciences, University of the Punjab, Quaid-e-Azam Campus, Lahore 54590, Pakistan.
Organizational Affiliation: