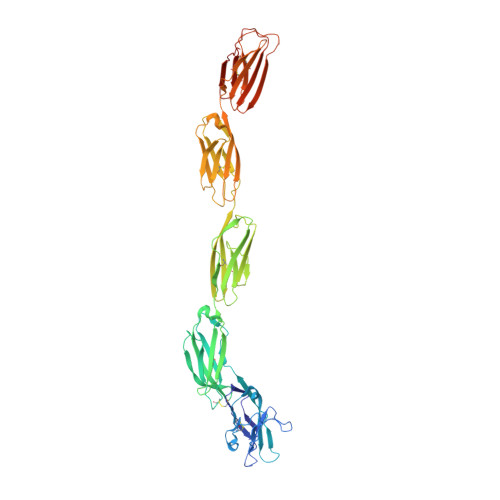Structural basis of myelin-associated glycoprotein adhesion and signalling.
Pronker, M.F., Lemstra, S., Snijder, J., Heck, A.J., Thies-Weesie, D.M., Pasterkamp, R.J., Janssen, B.J.(2016) Nat Commun 7: 13584-13584
- PubMed: 27922006 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms13584
- Primary Citation Related Structures:
5LF5, 5LFR, 5LFU, 5LFV - PubMed Abstract:
Myelin-associated glycoprotein (MAG) is a myelin-expressed cell-adhesion and bi-directional signalling molecule. MAG maintains the myelin-axon spacing by interacting with specific neuronal glycolipids (gangliosides), inhibits axon regeneration and controls myelin formation. The mechanisms underlying MAG adhesion and signalling are unresolved. We present crystal structures of the MAG full ectodomain, which reveal an extended conformation of five Ig domains and a homodimeric arrangement involving membrane-proximal domains Ig4 and Ig5. MAG-oligosaccharide complex structures and biophysical assays show how MAG engages axonal gangliosides at domain Ig1. Two post-translational modifications were identified-N-linked glycosylation at the dimerization interface and tryptophan C-mannosylation proximal to the ganglioside binding site-that appear to have regulatory functions. Structure-guided mutations and neurite outgrowth assays demonstrate MAG dimerization and carbohydrate recognition are essential for its regeneration-inhibiting properties. The combination of trans ganglioside binding and cis homodimerization explains how MAG maintains the myelin-axon spacing and provides a mechanism for MAG-mediated bi-directional signalling.
- Crystal and Structural Chemistry, Bijvoet Center for Biomolecular Research, Department of Chemistry, Faculty of Science, Utrecht University, Padualaan 8, 3584 CH Utrecht, The Netherlands.
Organizational Affiliation: