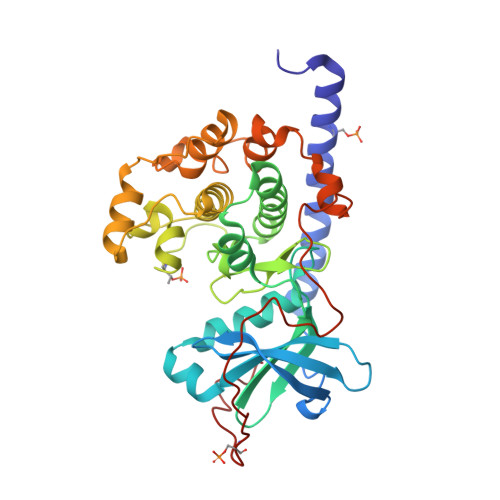
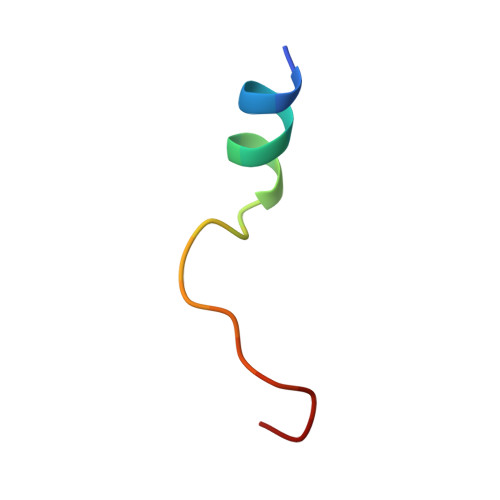
Cocrystal structure of cAMP-dependent Protein Kinase (PKA) in complex with different Fasudil-derivatives
Wienen-Schmidt, B., Jonker, H.R.A., Wulsdorf, T., Gerber, H.-D., Saxena, K., Kudlinzki, D., Sreeramulu, S., Parigi, G., Luchinat, C., Heine, A., Schwalbe, H., Klebe, G.To be published.