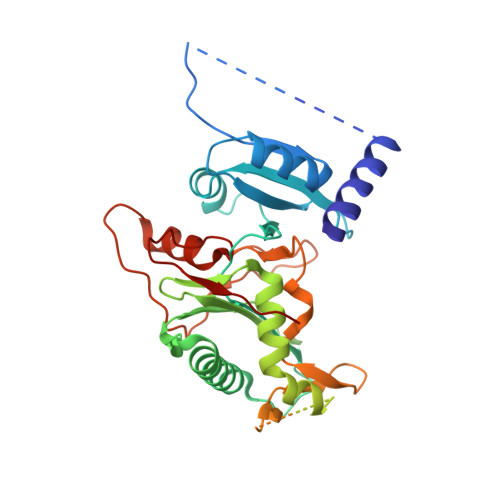
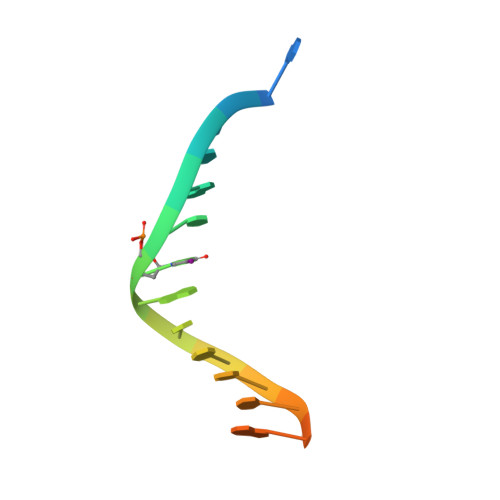
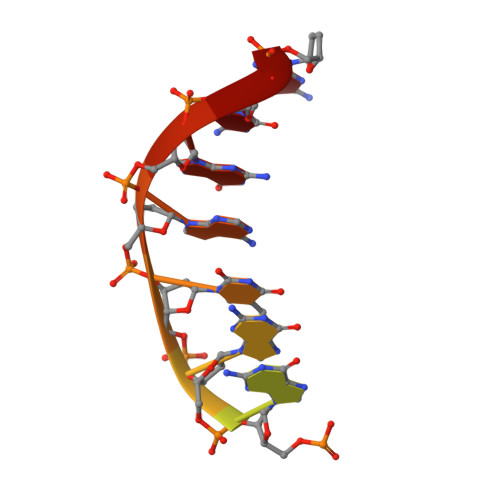
Structure and mechanism of human PrimPol, a DNA polymerase with primase activity.
Rechkoblit, O., Gupta, Y.K., Malik, R., Rajashankar, K.R., Johnson, R.E., Prakash, L., Prakash, S., Aggarwal, A.K.(2016) Sci Adv 2: e1601317-e1601317
- PubMed: 27819052 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.1601317
- Primary Citation Related Structures:
5L2X - PubMed Abstract:
PrimPol is a novel human enzyme that contains both DNA primase and DNA polymerase activities. We present the first structure of human PrimPol in ternary complex with a DNA template-primer and an incoming deoxynucleoside triphosphate (dNTP). The ability of PrimPol to function as a DNA primase stems from a simple but remarkable feature-almost complete lack of contacts to the DNA primer strand. This, in turn, allows two dNTPs to bind initiation and elongation sites on the enzyme for the formation of the first dinucleotide. PrimPol shows the ability to synthesize DNA opposite ultraviolet (UV) lesions; however, unexpectedly, the active-site cleft of the enzyme is constrained, which precludes the bypass of UV-induced DNA lesions by conventional translesion synthesis. Together, the structure addresses long-standing questions about how DNA primases actually initiate synthesis and how primase and polymerase activities combine in a single enzyme to carry out DNA synthesis.
- Department of Pharmacological Sciences, Icahn School of Medicine at Mount Sinai, Box 1677, 1425 Madison Avenue, New York, NY 10029, USA.
Organizational Affiliation: