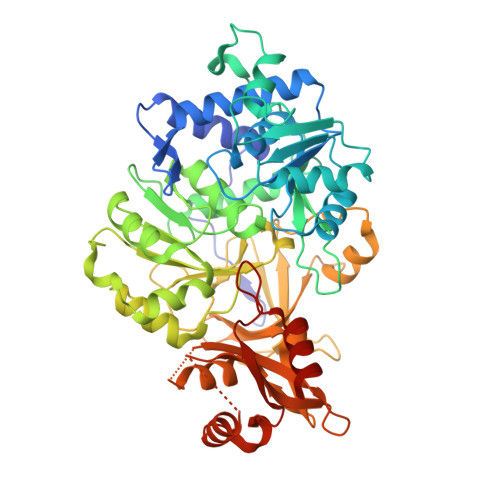
Cloning of the Orange Light-Producing Luciferase from Photinus scintillans-A New Proposal on how Bioluminescence Color is Determined.
Branchini, B.R., Southworth, T.L., Fontaine, D.M., Murtiashaw, M.H., McGurk, A., Talukder, M.H., Qureshi, R., Yetil, D., Sundlov, J.A., Gulick, A.M.(2017) Photochem Photobiol 93: 479-485
- PubMed: 27861940 Search on PubMed
- DOI: https://doi.org/10.1111/php.12671
- Primary Citation Related Structures:
5KYT - PubMed Abstract:
Unlike the enchanting yellow-green flashes of light produced on warm summer evenings by Photinus pyralis, the most common firefly species in North America, the orange lights of Photinus scintillans are infrequently observed. These Photinus species, and likely all bioluminescent beetles, use the same substrates beetle luciferin, ATP and oxygen to produce light. It is the structure of the particular luciferase enzyme that is the key to determining the color of the emitted light. We report here the molecular cloning of the P. scintillans luc gene and the expression and characterization of the corresponding novel recombinant luciferase enzyme. A comparison of the amino acid sequence with that of the highly similar P. pyralis enzyme and subsequent mutagenesis studies revealed that the single conservative amino acid change tyrosine to phenylalanine at position 255 accounted for the entire emission color difference. Additional mutagenesis and crystallographic studies were performed on a H-bond network, which includes the position 255 residue and five other stringently conserved beetle luciferase residues, that is proximal to the substrate/emitter binding site. The results are interpreted in the context of a speculative proposal that this network is key to the understanding of bioluminescence color determination.
- Department of Chemistry, Connecticut College, New London, CT.
Organizational Affiliation: