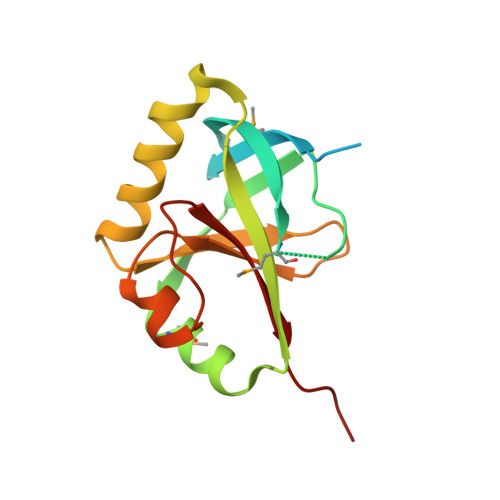
Crystal Structure of the SPOC Domain of the Arabidopsis Flowering Regulator FPA.
Zhang, Y., Rataj, K., Simpson, G.G., Tong, L.(2016) PLoS One 11: e0160694-e0160694
- PubMed: 27513867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0160694
- Primary Citation Related Structures:
5KXF - PubMed Abstract:
The Arabidopsis protein FPA controls flowering time by regulating the alternative 3'-end processing of the FLOWERING LOCUS (FLC) antisense RNA. FPA belongs to the split ends (SPEN) family of proteins, which contain N-terminal RNA recognition motifs (RRMs) and a SPEN paralog and ortholog C-terminal (SPOC) domain. The SPOC domain is highly conserved among FPA homologs in plants, but the conservation with the domain in other SPEN proteins is much lower. We have determined the crystal structure of Arabidopsis thaliana FPA SPOC domain at 2.7 Å resolution. The overall structure is similar to that of the SPOC domain in human SMRT/HDAC1 Associated Repressor Protein (SHARP), although there are also substantial conformational differences between them. Structural and sequence analyses identify a surface patch that is conserved among plant FPA homologs. Mutations of two residues in this surface patch did not disrupt FPA functions, suggesting that either the SPOC domain is not required for the role of FPA in regulating RNA 3'-end formation or the functions of the FPA SPOC domain cannot be disrupted by the combination of mutations, in contrast to observations with the SHARP SPOC domain.
- Department of Biological Sciences, Columbia University, New York, NY, 10027, United States of America.
Organizational Affiliation: