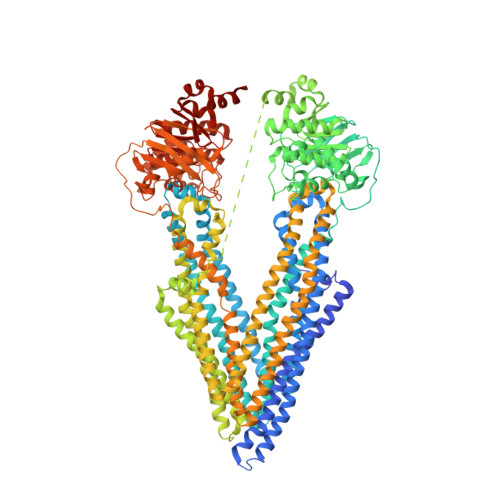
Structures of the Multidrug Transporter P-glycoprotein Reveal Asymmetric ATP Binding and the Mechanism of Polyspecificity.
Esser, L., Zhou, F., Pluchino, K.M., Shiloach, J., Ma, J., Tang, W.K., Gutierrez, C., Zhang, A., Shukla, S., Madigan, J.P., Zhou, T., Kwong, P.D., Ambudkar, S.V., Gottesman, M.M., Xia, D.(2017) J Biological Chem 292: 446-461
- PubMed: 27864369 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.755884
- Primary Citation Related Structures:
5KO2, 5KOY, 5KPD, 5KPI, 5KPJ - PubMed Abstract:
P-glycoprotein (P-gp) is a polyspecific ATP-dependent transporter linked to multidrug resistance in cancer; it plays important roles in determining the pharmacokinetics of many drugs. Understanding the structural basis of P-gp, substrate polyspecificity has been hampered by its intrinsic flexibility, which is facilitated by a 75-residue linker that connects the two halves of P-gp. Here we constructed a mutant murine P-gp with a shortened linker to facilitate structural determination. Despite dramatic reduction in rhodamine 123 and calcein-AM transport, the linker-shortened mutant P-gp possesses basal ATPase activity and binds ATP only in its N-terminal nucleotide-binding domain. Nine independently determined structures of wild type, the linker mutant, and a methylated P-gp at up to 3.3 Å resolution display significant movements of individual transmembrane domain helices, which correlated with the opening and closing motion of the two halves of P-gp. The open-and-close motion alters the surface topology of P-gp within the drug-binding pocket, providing a mechanistic explanation for the polyspecificity of P-gp in substrate interactions.
- From the Laboratory of Cell Biology, Center for Cancer Research, NCI.
Organizational Affiliation: