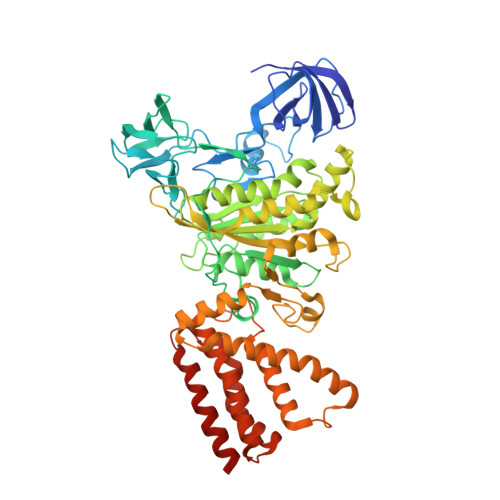
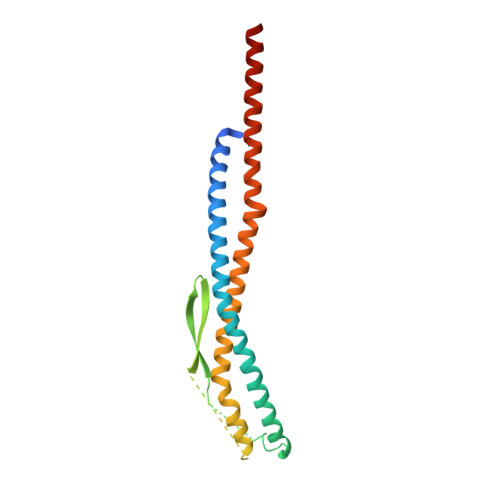
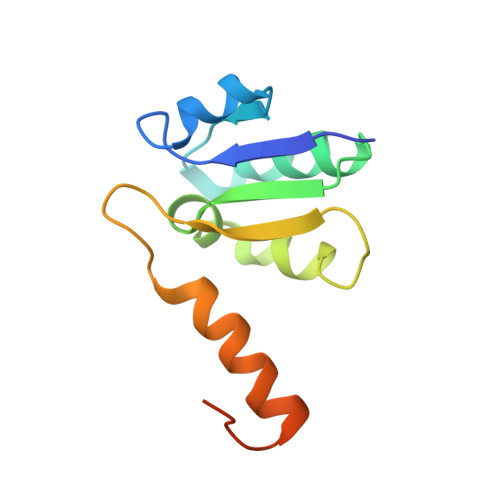
Crystal structures of the ATP-binding and ADP-release dwells of the V1 rotary motor
Suzuki, K., Mizutani, K., Maruyama, S., Shimono, K., Imai, F.L., Muneyuki, E., Kakinuma, Y., Ishizuka-Katsura, Y., Shirouzu, M., Yokoyama, S., Yamato, I., Murata, T.(2016) Nat Commun 7: 13235-13235
- PubMed: 27807367 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms13235
- Primary Citation Related Structures:
5KNB, 5KNC, 5KND - PubMed Abstract:
V 1 -ATPases are highly conserved ATP-driven rotary molecular motors found in various membrane systems. We recently reported the crystal structures for the Enterococcus hirae A 3 B 3 DF (V 1 ) complex, corresponding to the catalytic dwell state waiting for ATP hydrolysis. Here we present the crystal structures for two other dwell states obtained by soaking nucleotide-free V 1 crystals in ADP. In the presence of 20 μM ADP, two ADP molecules bind to two of three binding sites and cooperatively induce conformational changes of the third site to an ATP-binding mode, corresponding to the ATP-binding dwell. In the presence of 2 mM ADP, all nucleotide-binding sites are occupied by ADP to induce conformational changes corresponding to the ADP-release dwell. Based on these and previous findings, we propose a V 1 -ATPase rotational mechanism model.
- Department of Chemistry, Graduate School of Science, Chiba University, 1-33 Yayoi-cho, Inage, Chiba 263-8522, Japan.
Organizational Affiliation: