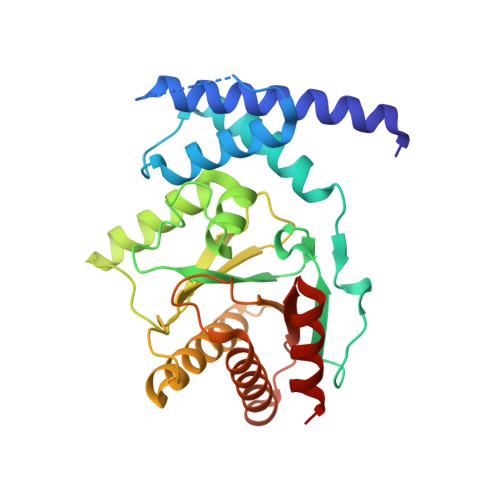
Structure of the sirtuin-linked macrodomain SAV0325 from Staphylococcus aureus.
Appel, C.D., Feld, G.K., Wallace, B.D., Williams, R.S.(2016) Protein Sci 25: 1682-1691
- PubMed: 27345688
- DOI: https://doi.org/10.1002/pro.2974
- Primary Citation Related Structures:
5KIV - PubMed Abstract:
Cells use the post-translational modification ADP-ribosylation to control a host of biological activities. In some pathogenic bacteria, an operon-encoded mono-ADP-ribosylation cycle mediates response to host-induced oxidative stress. In this system, reversible mono ADP-ribosylation of a lipoylated target protein represses oxidative stress response. An NAD(+) -dependent sirtuin catalyzes the single ADP-ribose (ADPr) addition, while a linked macrodomain-containing protein removes the ADPr. Here we report the crystal structure of the sitruin-linked macrodomain protein from Staphylococcus aureus, SauMacro (also known as SAV0325) to 1.75-Å resolution. The monomeric SauMacro bears a previously unidentified Zn(2+) -binding site that putatively aids in substrate recognition and catalysis. An amino-terminal three-helix bundle motif unique to this class of macrodomain proteins provides a structural scaffold for the Zn(2+) site. Structural features of the enzyme further indicate a cleft proximal to the Zn(2+) binding site appears well suited for ADPr binding, while a deep hydrophobic channel in the protein core is suitable for binding the lipoate of the lipoylated protein target.
- Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences, US National Institutes of Health, Department of Health and Human Services, Research Triangle Park, North Carolina, 27709.
Organizational Affiliation: