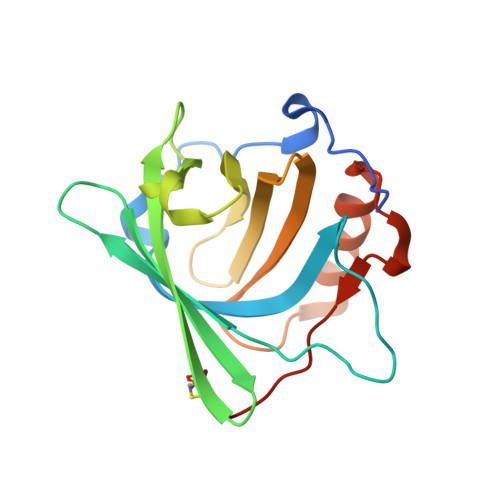
Chelation and stabilization of berkelium in oxidation state +IV.
Deblonde, G.J., Sturzbecher-Hoehne, M., Rupert, P.B., An, D.D., Illy, M.C., Ralston, C.Y., Brabec, J., de Jong, W.A., Strong, R.K., Abergel, R.J.(2017) Nat Chem 9: 843-849
- PubMed: 28837177 Search on PubMed
- DOI: https://doi.org/10.1038/nchem.2759
- Primary Citation Related Structures:
5KIC - PubMed Abstract:
Berkelium (Bk) has been predicted to be the only transplutonium element able to exhibit both +III and +IV oxidation states in solution, but evidence of a stable oxidized Bk chelate has so far remained elusive. Here we describe the stabilization of the heaviest 4+ ion of the periodic table, under mild aqueous conditions, using a siderophore derivative. The resulting Bk(IV) complex exhibits luminescence via sensitization through an intramolecular antenna effect. This neutral Bk(IV) coordination compound is not sequestered by the protein siderocalin-a mammalian metal transporter-in contrast to the negatively charged species obtained with neighbouring trivalent actinides americium, curium and californium (Cf). The corresponding Cf(III)-ligand-protein ternary adduct was characterized by X-ray diffraction analysis. Combined with theoretical predictions, these data add significant insight to the field of transplutonium chemistry, and may lead to innovative Bk separation and purification processes.
- Chemical Sciences Division, Lawrence Berkeley National Laboratory, Berkeley, California 94720, USA.
Organizational Affiliation: