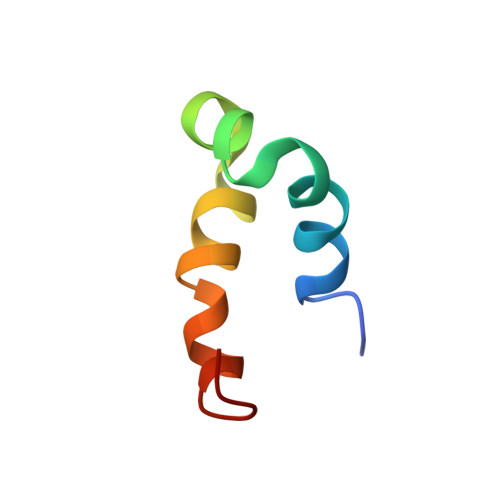
Solution Structures of Phenol-Soluble Modulins alpha 1, alpha 3, and beta 2, Virulence Factors from Staphylococcus aureus.
Towle, K.M., Lohans, C.T., Miskolzie, M., Acedo, J.Z., van Belkum, M.J., Vederas, J.C.(2016) Biochemistry 55: 4798-4806
- PubMed: 27525453
- DOI: https://doi.org/10.1021/acs.biochem.6b00615
- Primary Citation Related Structures:
5KGY, 5KGZ, 5KHB - PubMed Abstract:
Phenol-soluble modulins (PSMs) are peptide virulence factors produced by staphylococci. These peptides contribute to the overall pathogenicity of these bacteria, eliciting multiple immune responses from host cells. Many of the α-type PSMs exhibit cytolytic properties and are able to lyse particular eukaryotic cells, including erythrocytes, neutrophils, and leukocytes. In addition, they also appear to contribute to the protection of the bacterial cell from the host immune response through biofilm formation and detachment. In this study, three of these peptide toxins, PSMs α1, α3, and β2, normally produced by Staphylococcus aureus, have been synthesized using solid-supported peptide synthesis (SPPS) (PSMα1 and PSMα3) or made by heterologous expression in Escherichia coli (PSMβ2). Their three-dimensional structures were elucidated using nuclear magnetic resonance spectroscopy. PSMα1 and PSMα3 each consist of a single amphipathic helix with a slight bend near the N- and C-termini, respectively. PSMβ2 contains three amphipathic helices, which fold to produce a "v-like" shape between α-helix 2 and α-helix 3, with α-helix 1 folded over such that it is perpendicular to α-helix 3. The availability of three-dimensional structures permits spatial analysis of features and residues proposed to control the biological activity of these peptide toxins.
- Department of Chemistry, University of Alberta , Edmonton, Alberta, Canada T6G 2G2.
Organizational Affiliation: