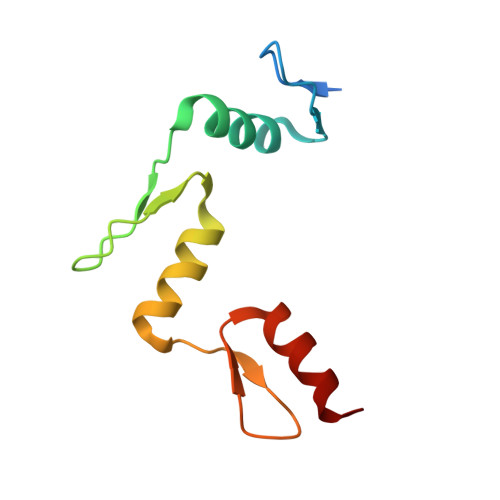
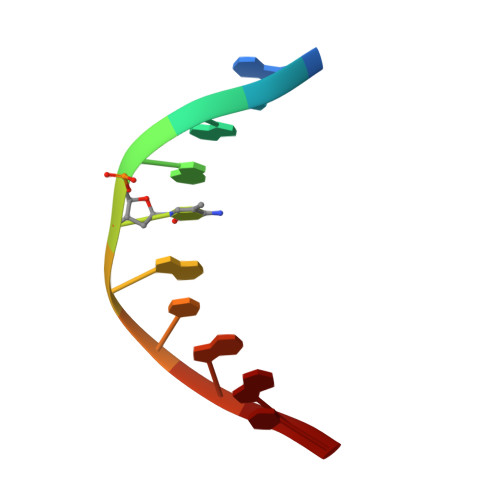
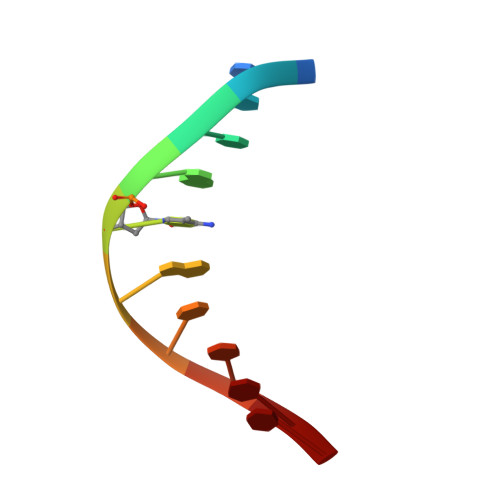
Distinctive Klf4 mutants determine preference for DNA methylation status.
Hashimoto, H., Wang, D., Steves, A.N., Jin, P., Blumenthal, R.M., Zhang, X., Cheng, X.(2016) Nucleic Acids Res 44: 10177-10185
- PubMed: 27596594 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkw774
- Primary Citation Related Structures:
5KE6, 5KE7, 5KE8, 5KE9, 5KEA, 5KEB - PubMed Abstract:
Reprogramming of mammalian genome methylation is critically important but poorly understood. Klf4, a transcription factor directing reprogramming, contains a DNA binding domain with three consecutive C2H2 zinc fingers. Klf4 recognizes CpG or TpG within a specific sequence. Mouse Klf4 DNA binding domain has roughly equal affinity for methylated CpG or TpG, and slightly lower affinity for unmodified CpG. The structural basis for this key preference is unclear, though the side chain of Glu446 is known to contact the methyl group of 5-methylcytosine (5mC) or thymine (5-methyluracil). We examined the role of Glu446 by mutagenesis. Substituting Glu446 with aspartate (E446D) resulted in preference for unmodified cytosine, due to decreased affinity for 5mC. In contrast, substituting Glu446 with proline (E446P) increased affinity for 5mC by two orders of magnitude. Structural analysis revealed hydrophobic interaction between the proline's aliphatic cyclic structure and the 5-methyl group of the pyrimidine (5mC or T). As in wild-type Klf4 (E446), the proline at position 446 does not interact directly with either the 5mC N4 nitrogen or the thymine O4 oxygen. In contrast, the unmethylated cytosine's exocyclic N4 amino group (NH 2 ) and its ring carbon C5 atom hydrogen bond directly with the aspartate carboxylate of the E446D variant. Both of these interactions would provide a preference for cytosine over thymine, and the latter one could explain the E446D preference for unmethylated cytosine. Finally, we evaluated the ability of these Klf4 mutants to regulate transcription of methylated and unmethylated promoters in a luciferase reporter assay.
- Department of Biochemistry, Emory University, Atlanta, GA 30322, USA.
Organizational Affiliation: