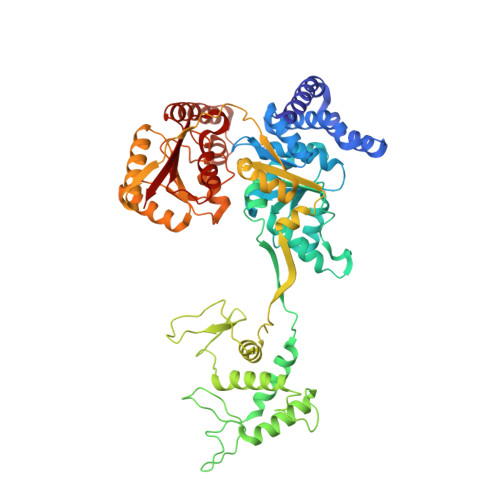
An alternate mode of oligomerization for E. coli SecA.
Yazdi, A.K., Vezina, G.C., Shilton, B.H.(2017) Sci Rep 7: 11747-11747
- PubMed: 28924213
- DOI: https://doi.org/10.1038/s41598-017-11648-5
- Primary Citation Related Structures:
5K9T - PubMed Abstract:
SecA is the ATPase of preprotein translocase. SecA is a dimer in solution and changes in its oligomeric state may function in preprotein translocation. The SecA-N68 construct, in which the C-terminal helical domains of SecA are deleted, was used to investigate the mechanism of SecA oligomerization. SecA-N68 is in equilibrium between monomers, dimers, and tetramers. Subunit interactions in the SecA-N68 tetramer are mediated entirely by unstructured regions at its N- and C-termini: when the termini are deleted to yield SecA-N68∆NC, the construct is completely monomeric. This monomeric construct yielded crystals diffracting to 2.6 Å that were used to solve the structure of SecA-N68, including the "preprotein crosslinking domain" (PPXD) that was missing from previous E. coli SecA structures. The SecA-N68 structure was combined with small angle X-ray scattering (SAXS) data to construct a model of the SecA-N68 tetramer that is consistent with the essential roles of the extreme N- and C-termini in oligomerization. This mode of oligomerization, which depends on binding of the extreme N-terminus to the DEAD motor domains, NBD1 and NBD2, was used to model a novel parallel and flexible SecA solution dimer that agrees well with SAXS data.
- Department of Biochemistry, University of Western Ontario, London, Ontario, N6A 5C1, Canada.
Organizational Affiliation: