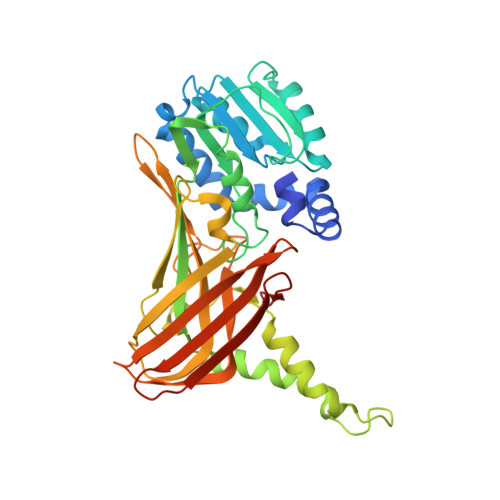
Structural studies of protein arginine methyltransferase 2 reveal its interactions with potential substrates and inhibitors.
Cura, V., Marechal, N., Troffer-Charlier, N., Strub, J.M., van Haren, M.J., Martin, N.I., Cianferani, S., Bonnefond, L., Cavarelli, J.(2017) FEBS J 284: 77-96
- PubMed: 27879050 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13953
- Primary Citation Related Structures:
5FUB, 5FUL, 5FWA, 5FWD, 5G02, 5JMQ, 5K8V - PubMed Abstract:
PRMT2 is the less-characterized member of the protein arginine methyltransferase family in terms of structure, activity, and cellular functions. PRMT2 is a modular protein containing a catalytic Ado-Met-binding domain and unique Src homology 3 domain that binds proteins with proline-rich motifs. PRMT2 is involved in a variety of cellular processes and has diverse roles in transcriptional regulation through different mechanisms depending on its binding partners. PRMT2 has been demonstrated to have weak methyltransferase activity on a histone H4 substrate, but its optimal substrates have not yet been identified. To obtain insights into the function and activity of PRMT2, we solve several crystal structures of PRMT2 from two homologs (zebrafish and mouse) in complex with either the methylation product S-adenosyl-L-homocysteine or other compounds including the first synthetic PRMT2 inhibitor (Cp1) studied so far. We reveal that the N-terminal-containing SH3 module is disordered in the full-length crystal structures, and highlights idiosyncratic features of the PRMT2 active site. We identify a new nonhistone protein substrate belonging to the serine-/arginine-rich protein family which interacts with PRMT2 and we characterize six methylation sites by mass spectrometry. To better understand structural basis for Cp1 binding, we also solve the structure of the complex PRMT4:Cp1. We compare the inhibitor-protein interactions occurring in the PRMT2 and PRMT4 complex crystal structures and show that this compound inhibits efficiently PRMT2. These results are a first step toward a better understanding of PRMT2 substrate recognition and may accelerate the development of structure-based drug design of PRMT2 inhibitors. All coordinates and structure factors have been deposited in the Protein Data Bank: zPRMT2 1-408 -SFG = 5g02; zPRMT2 73-408 -SAH = 5fub; mPRMT2 1-445 -SAH = 5ful; mPRMT2 1-445 -Cp1 = 5fwa, mCARM1 130-487 -Cp1 = 5k8v.
- Department of Integrated Structural Biology, Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), CNRS UMR7104, INSERM U596, Université de Strasbourg, Illkirch, France.
Organizational Affiliation: