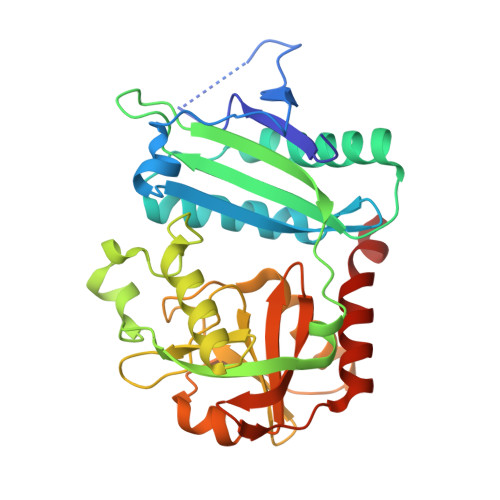
Discovery and structural characterisation of new fold type IV-transaminases exemplify the diversity of this enzyme fold.
Pavkov-Keller, T., Strohmeier, G.A., Diepold, M., Peeters, W., Smeets, N., Schurmann, M., Gruber, K., Schwab, H., Steiner, K.(2016) Sci Rep 6: 38183-38183
- PubMed: 27905516
- DOI: https://doi.org/10.1038/srep38183
- Primary Citation Related Structures:
5K3W - PubMed Abstract:
Transaminases are useful biocatalysts for the production of amino acids and chiral amines as intermediates for a broad range of drugs and fine chemicals. Here, we describe the discovery and characterisation of new transaminases from microorganisms which were enriched in selective media containing (R)-amines as sole nitrogen source. While most of the candidate proteins were clearly assigned to known subgroups of the fold IV family of PLP-dependent enzymes by sequence analysis and characterisation of their substrate specificity, some of them did not fit to any of these groups. The structure of one of these enzymes from Curtobacterium pusillum, which can convert d-amino acids and various (R)-amines with high enantioselectivity, was solved at a resolution of 2.4 Å. It shows significant differences especially in the active site compared to other transaminases of the fold IV family and thus indicates the existence of a new subgroup within this family. Although the discovered transaminases were not able to convert ketones in a reasonable time frame, overall, the enrichment-based approach was successful, as we identified two amine transaminases, which convert (R)-amines with high enantioselectivity, and can be used for a kinetic resolution of 1-phenylethylamine and analogues to obtain the (S)-amines with e.e.s >99%.
- acib, Austrian Centre of Industrial Biotechnology GmbH, 8010 Graz, Austria.
Organizational Affiliation: