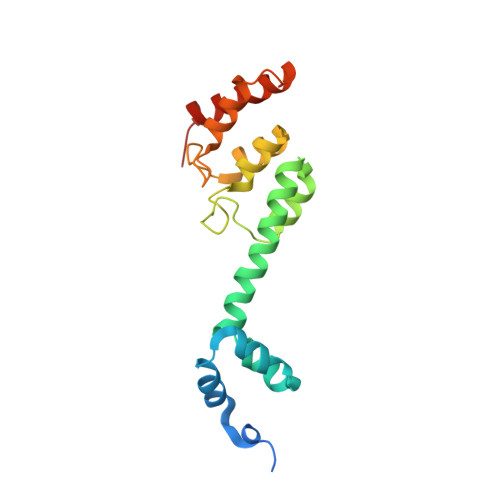
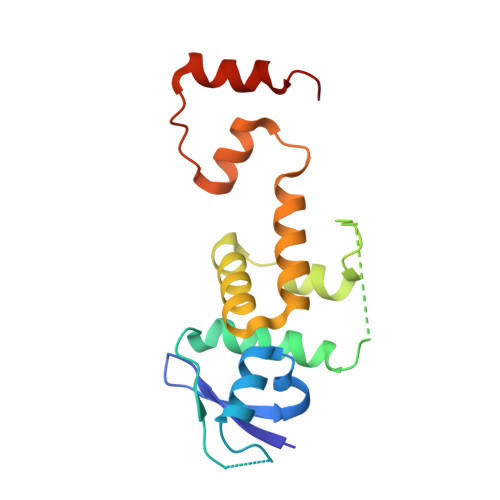
Structural Mimicry by a Bacterial F Box Effector Hijacks the Host Ubiquitin-Proteasome System.
Wong, K., Perpich, J.D., Kozlov, G., Cygler, M., Abu Kwaik, Y., Gehring, K.(2017) Structure 25: 376-383
- PubMed: 28111017 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.12.015
- Primary Citation Related Structures:
5K34, 5K35 - PubMed Abstract:
Ankyrin B (AnkB/LegAU13) is a translocated F box effector essential for the intracellular replication of the pathogen Legionella pneumophila. AnkB co-opts a host ubiquitin ligase to decorate the pathogen-containing vacuole with K 48 -linked polyubiquitinated proteins and degrade host proteins as a source of energy. Here, we report that AnkB commandeers the host ubiquitin-proteasome system through mimicry of two eukaryotic protein domains. Using X-ray crystallography, we determined the 3D structure of AnkB in complex with Skp1, a component of the human SCF ubiquitination ligase. The structure confirms that AnkB contains an N-terminal F box similar to Skp2 and a C-terminal substrate-binding domain similar to eukaryotic ankyrin repeats. We identified crucial amino acids in the substrate-binding domain of AnkB and showed them to be essential for the function of AnkB in L. pneumophila intracellular proliferation. The study reveals how Legionella uses molecular mimicry to manipulate the host ubiquitination pathway and proliferate intracellularly.
- Department of Biochemistry and Groupe de recherche axé sur la structure des protéines, McGill University, Montreal, QC H3G 0B1, Canada.
Organizational Affiliation: