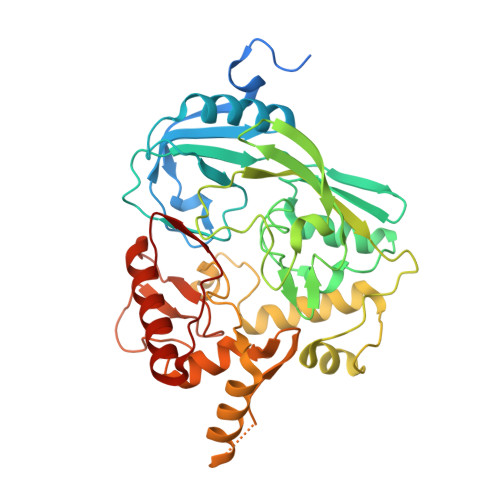
Crystal structure of UDP-N-acetylglucosamine-enolpyruvate reductase (MurB) from Mycobacterium tuberculosis
Eniyan, K., Dharavath, S., Vijayan, R., Bajpai, U., Gourinath, S.(2017) Biochim Biophys Acta 1866: 397-406
- PubMed: 29203374 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2017.11.013
- Primary Citation Related Structures:
5JZX - PubMed Abstract:
The biosynthesis of UDP-N-acetylmuramic acid (UDP-MurNAc) by reduction of UDP-N-acetylglucosamine-enolpyruvate (UDP-GlcNAc-EP) in an NADPH and FAD-dependent reaction in bacteria is one of the key steps in peptidoglycan biosynthesis catalyzed by UDP-N-acetylglucosamine-enolpyruvate reductase (MurB). Here, we present the crystal structure of Mycobacterium tuberculosis MurB (MtbMurB) with FAD as the prosthetic group at 2.0Å resolution. There are six molecules in asymmetric unit in the form of dimers. Each protomer can be subdivided into three domains and the prosthetic group, FAD is bound in the active site between domain I and domain II. Comparison of MtbMurB structure with the structures of the Escherichia coli MurB (in complex with UDP-GlcNAc-EP) and Pseudomonas aeruginosa MurB (in complex with NADPH) showed all three structures share similar domain architecture and residues in the active site. The nicotinamide and the enol pyruvyl moieties are well aligned upon superimposition, both positioned in suitable position for hydride transfer to and from FAD. The comparison studies and MD simulations demonstrate that the two lobes of domain-III become more flexible. The substrates (NADPH and UDP-GlcNAc-EP) binding responsible for open conformation of MurB, suggesting that NADPH and UDP-GlcNAc-EP interactions are conformationally stable. Our findings provide a detail mechanism about the closed to open state by binding of NADPH and UDP-GlcNAc-EP induces the conformational changes of MurB structure that may trigger the MurB catalytic reaction.
- Department of Biomedical Science, Acharya Narendra Dev College, University of Delhi, New Delhi, India.
Organizational Affiliation: