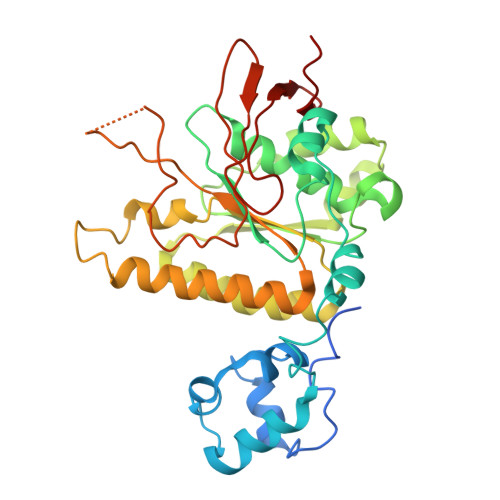
High-resolution structure of the presynaptic RAD51 filament on single-stranded DNA by electron cryo-microscopy.
Short, J.M., Liu, Y., Chen, S., Soni, N., Madhusudhan, M.S., Shivji, M.K., Venkitaraman, A.R.(2016) Nucleic Acids Res 44: 9017-9030
- PubMed: 27596592
- DOI: https://doi.org/10.1093/nar/gkw783
- Primary Citation of Related Structures:
5JZC, 5NP7 - PubMed Abstract:
Homologous DNA recombination (HR) by the RAD51 recombinase enables error-free DNA break repair. To execute HR, RAD51 first forms a presynaptic filament on single-stranded (ss) DNA, which catalyses pairing with homologous double-stranded (ds) DNA. Here, we report a structure for the presynaptic human RAD51 filament at 3.5-5.0Å resolution using electron cryo-microscopy. RAD51 encases ssDNA in a helical filament of 103Å pitch, comprising 6.4 protomers per turn, with a rise of 16.1Å and a twist of 56.2°. Inter-protomer distance correlates with rotation of an α-helical region in the core catalytic domain that is juxtaposed to ssDNA, suggesting how the RAD51-DNA interaction modulates protomer spacing and filament pitch. We map Fanconi anaemia-like disease-associated RAD51 mutations, clarifying potential phenotypes. We predict binding sites on the presynaptic filament for two modules present in each BRC repeat of the BRCA2 tumour suppressor, a critical HR mediator. Structural modelling suggests that changes in filament pitch mask or expose one binding site with filament-inhibitory potential, rationalizing the paradoxical ability of the BRC repeats to either stabilize or inhibit filament formation at different steps during HR. Collectively, our findings provide fresh insight into the structural mechanism of HR and its dysregulation in human disease.
- Medical Research Council Cancer Unit, University of Cambridge, Hills Road, Cambridge CB2 0XZ, UK.
Organizational Affiliation: