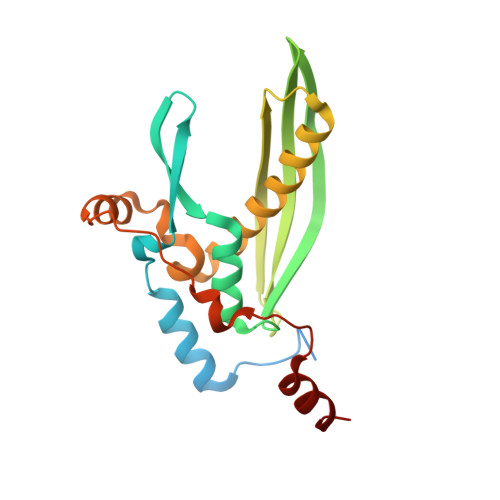
Structure of the human DNA-repair protein RAD52 containing surface mutations.
Saotome, M., Saito, K., Onodera, K., Kurumizaka, H., Kagawa, W.(2016) Acta Crystallogr F Struct Biol Commun 72: 598-603
- PubMed: 27487923 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X1601027X
- Primary Citation Related Structures:
5JRB - PubMed Abstract:
The Rad52 protein is a eukaryotic single-strand DNA-annealing protein that is involved in the homologous recombinational repair of DNA double-strand breaks. The isolated N-terminal half of the human RAD52 protein (RAD52(1-212)) forms an undecameric ring structure with a surface that is mostly positively charged. In the present study, it was found that RAD52(1-212) containing alanine mutations of the charged surface residues (Lys102, Lys133 and Glu202) is highly amenable to crystallization. The structure of the mutant RAD52(1-212) was solved at 2.4 Å resolution. The structure revealed an association between the symmetry-related RAD52(1-212) rings, in which a partially unfolded, C-terminal region of RAD52 extended into the DNA-binding groove of the neighbouring ring in the crystal. The alanine mutations probably reduced the surface entropy of the RAD52(1-212) ring and stabilized the ring-ring association observed in the crystal.
- Department of Interdisciplinary Science and Engineering, Program in Chemistry and Life Science, School of Science and Engineering, Meisei University, 2-1-1 Hodokubo, Hino-shi, Tokyo 191-8506, Japan.
Organizational Affiliation: