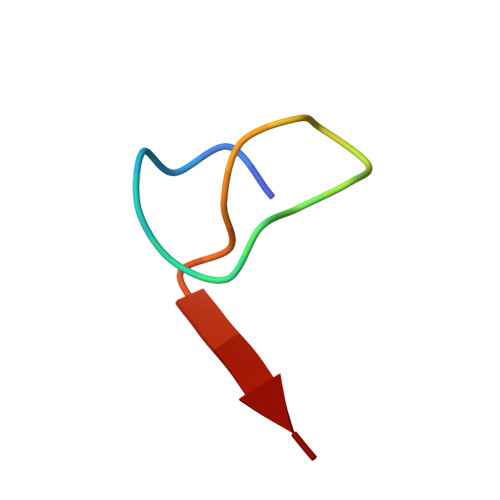
Structure and Mechanism of the Sphingopyxin I Lasso Peptide Isopeptidase.
Fage, C.D., Hegemann, J.D., Nebel, A.J., Steinbach, R.M., Zhu, S., Linne, U., Harms, K., Bange, G., Marahiel, M.A.(2016) Angew Chem Int Ed Engl 55: 12717-12721
- PubMed: 27611791
- DOI: https://doi.org/10.1002/anie.201605232
- Primary Citation Related Structures:
5JQF, 5JRK, 5JRL - PubMed Abstract:
Lasso peptides are natural products that assume a unique lariat knot topology. Lasso peptide isopeptidases (IsoPs) eliminate this topology through isopeptide bond cleavage. To probe how these enzymes distinguish between substrates and hydrolyze only isopeptide bonds, we examined the structure and mechanism of a previously uncharacterized IsoP from the proteobacterium Sphingopyxis alaskensis RB2256 (SpI-IsoP). We demonstrate that SpI-IsoP efficiently and specifically linearizes the lasso peptide sphingopyxin I (SpI) and variants thereof. We also present crystal structures of SpI and SpI-IsoP, revealing a threaded topology for the former and a prolyl oligopeptidase (POP)-like fold for the latter. Subsequent structure-guided mutational analysis allowed us to propose roles for active-site residues. Our study sheds light on lasso peptide catabolism and expands the engineering potential of these fascinating molecules.
- Fachbereich Chemie, Fachgebiet Biochemie und LOEWE-Zentrum für Synthetische Mikrobiologie, Philipps-Universität Marburg, Hans-Meerwein-Strasse 4, 35032, Marburg, Germany.
Organizational Affiliation: