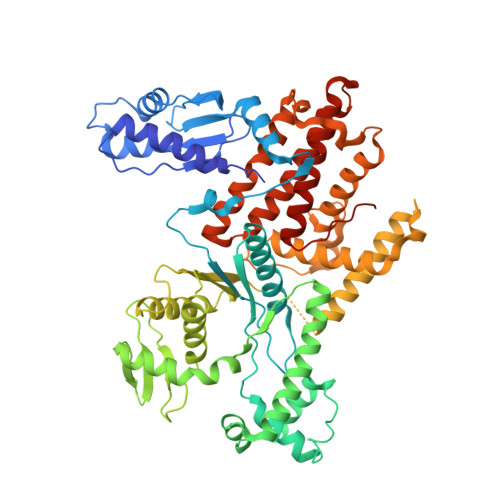
Dimerization of Arginyl-tRNA Synthetase by Free Heme Drives Its Inactivation in Plasmodium falciparum
Jain, V., Yogavel, M., Sharma, A.(2016) Structure 24: 1476-1487
- PubMed: 27502052
- DOI: https://doi.org/10.1016/j.str.2016.06.018
- Primary Citation Related Structures:
5JLD - PubMed Abstract:
Excess cellular heme is toxic, and malaria parasites regulate its levels during hemoglobin digestion. Aminoacyl-tRNA synthetases are ubiquitous enzymes, and of these, arginyl-tRNA synthetase (RRS) is unique as its enzymatic product of charged tRNA is required for protein synthesis and degradation. We show that Plasmodium falciparum arginyl-tRNA synthetase (PfRRS) is an active, cytosolic, and monomeric enzyme. Its high-resolution crystal structure highlights critical structural differences with the human enzyme. We further show that hemin binds to and inhibits the aminoacylation activity of PfRRS. Hemin induces a dimeric form of PfRRS that is thus rendered enzymatically dead as it is unable to recognize its cognate tRNA(arg). Excessive hemin in chloroquine-treated malaria parasites results in significantly reduced charged tRNA(arg) levels, thus suggesting deceleration of protein synthesis. These data together suggest that the inhibition of Plasmodium falciparum arginyl-tRNA synthetase can now be synergized with existing antimalarials for more potent drug cocktails against malaria parasites.
- Molecular Medicine Group, International Centre for Genetic Engineering and Biotechnology (ICGEB), Aruna Asaf Ali Road, New Delhi 110067, India.
Organizational Affiliation: