Structural Basis for Receptor-Mediated Selective Autophagy of Aminopeptidase I Aggregates
Yamasaki, A., Watanabe, Y., Adachi, W., Suzuki, K., Matoba, K., Kirisako, H., Kumeta, H., Nakatogawa, H., Ohsumi, Y., Inagaki, F., Noda, N.N.(2016) Cell Rep 16: 19-27
- PubMed: 27320913
- DOI: https://doi.org/10.1016/j.celrep.2016.05.066
- Primary Citation Related Structures:
5JGE, 5JGF, 5JH9, 5JHC - PubMed Abstract:
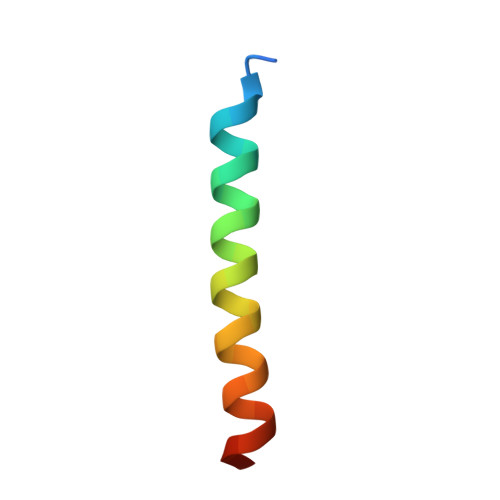
Selective autophagy mediates the degradation of various cargoes, including protein aggregates and organelles, thereby contributing to cellular homeostasis. Cargo receptors ensure selectivity by tethering specific cargo to lipidated Atg8 at the isolation membrane. However, little is known about the structural requirements underlying receptor-mediated cargo recognition. Here, we report structural, biochemical, and cell biological analysis of the major selective cargo protein in budding yeast, aminopeptidase I (Ape1), and its complex with the receptor Atg19. The Ape1 propeptide has a trimeric coiled-coil structure, which tethers dodecameric Ape1 bodies together to form large aggregates. Atg19 disassembles the propeptide trimer and forms a 2:1 heterotrimer, which not only blankets the Ape1 aggregates but also regulates their size. These receptor activities may promote elongation of the isolation membrane along the aggregate surface, enabling sequestration of the cargo with high specificity.
- Institute of Microbial Chemistry (BIKAKEN), Tokyo 141-0021, Japan.
Organizational Affiliation: