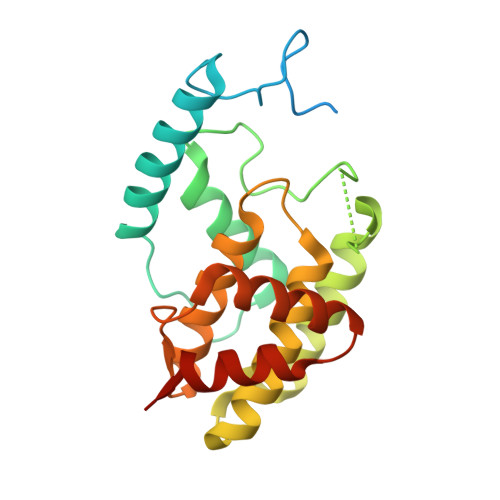
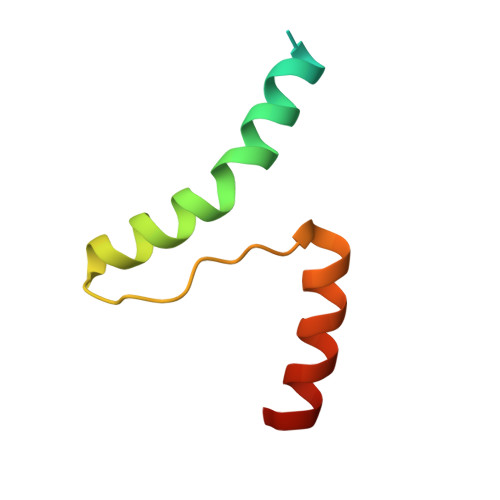
Crystal Structure of the Escherichia coli Fic Toxin-Like Protein in Complex with Its Cognate Antitoxin.
Stanger, F.V., Harms, A., Dehio, C., Schirmer, T.(2016) PLoS One 11: e0163654-e0163654
- PubMed: 27657533 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0163654
- Primary Citation Related Structures:
5JFF, 5JFZ - PubMed Abstract:
FIC domain proteins mediate post-translational modifications of target proteins, which typically results in their inactivation. Depending on the conservation of crucial active site residues, the FIC fold serves as structural scaffold for various enzymatic activities, mostly target adenylylation. The founding member of the vast Fic protein family, EcFicT, was identified in Escherichia coli some time ago. The G55R point mutant of EcFicT displays the "filamentation induced by cAMP" (Fic) phenotype at high 3',5'-cyclic adenosine monophosphate (cAMP) concentrations and elevated temperature, but the underlying molecular mechanism and any putative biochemical activity of EcFicT have remained unknown. EcFicT belongs to class I Fic toxin proteins that are encoded together with a small inhibitory protein (antitoxin), named EcFicA in E. coli. Here, we report the crystal structures of two mutant EcFicT/EcFicA complexes (EcFicTG55RA and EcFicTAE28G) both showing close resemblance with the structure of the AMP-transferase VbhT from Bartonella schoenbuchensis in complex with its cognate antitoxin VbhA. However, crucial differences in the active site of EcFicT compared to VbhT and other AMP-transferases rationalize the lack of evidence for adenylylation activity. Comprehensive bioinformatic analysis suggests that EcFicT has evolved from canonical AMP-transferases and has acquired a conserved binding site for a yet to be discovered novel substrate. The G55R mutation has no effect on structure or thermal stability of EcFicT, such that the molecular basis for its associated Fic phenotype remains elusive. We anticipate that this structure will inspire further bioinformatic and experimental analyses in order to characterize the enzymatic activity of EcFicT and help revealing its physiological role.
- Focal Area Structural Biology and Biophysics, Biozentrum, University of Basel, Basel, Switzerland.
Organizational Affiliation: