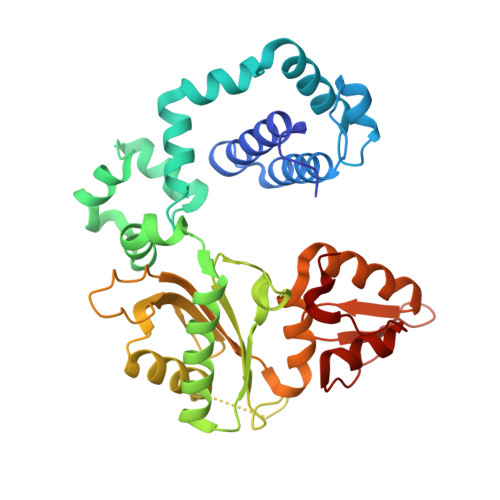
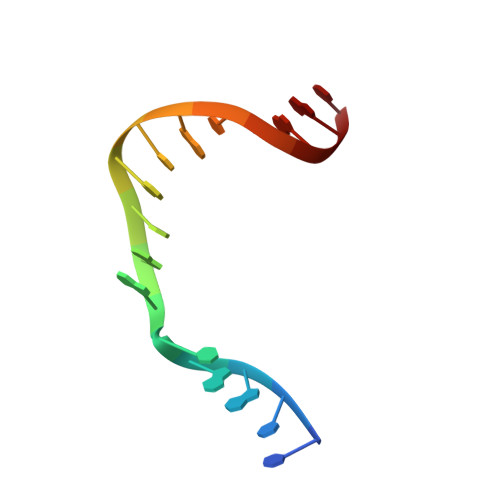
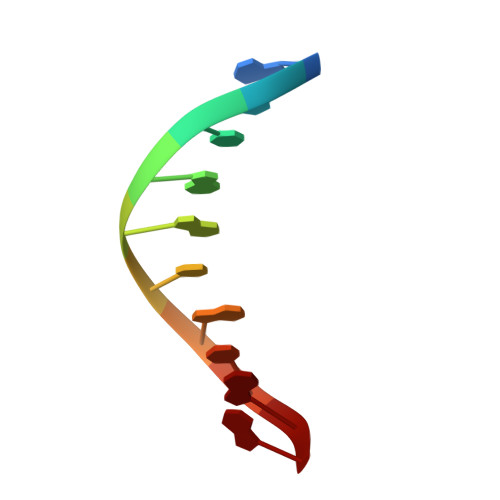
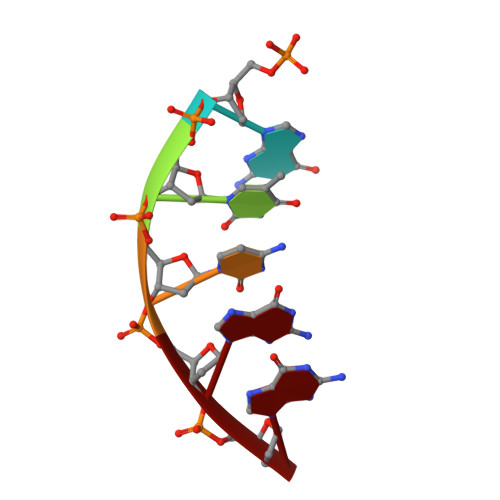
Structures of DNA Polymerase Mispaired DNA Termini Transitioning to Pre-catalytic Complexes Support an Induced-Fit Fidelity Mechanism.
Batra, V.K., Beard, W.A., Pedersen, L.C., Wilson, S.H.(2016) Structure 24: 1863-1875
- PubMed: 27642161 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.08.006
- Primary Citation Related Structures:
5J0O, 5J0P, 5J0Q, 5J0R, 5J0S, 5J0T, 5J0U, 5J0W, 5J0X, 5J0Y, 5J29, 5J2A, 5J2B, 5J2C, 5J2D, 5J2E, 5J2F, 5J2G, 5J2H, 5J2I, 5J2J, 5J2K, 5TZV - PubMed Abstract:
High-fidelity DNA synthesis requires that polymerases display a strong preference for right nucleotide insertion. When the wrong nucleotide is inserted, the polymerase deters extension from the mismatched DNA terminus. Twenty-three crystallographic structures of DNA polymerase β with terminal template-primer mismatches were determined as binary DNA and ternary pre-catalytic substrate complexes. These structures indicate that the mismatched termini adopt various distorted conformations that attempt to satisfy stacking and hydrogen-bonding interactions. The binary complex structures indicate an induced strain in the mismatched template nucleotide. Addition of a non-hydrolyzable incoming nucleotide stabilizes the templating nucleotide with concomitant strain in the primer terminus. Several dead-end ternary complex structures suggest that DNA synthesis might occur as the enzyme transitions from an open to a closed complex. The structures are consistent with an induced-fit mechanism where a mismatched terminus is misaligned relative to the correct incoming nucleotide to deter or delay further DNA synthesis.
- Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, NC 27709-12233, USA.
Organizational Affiliation: