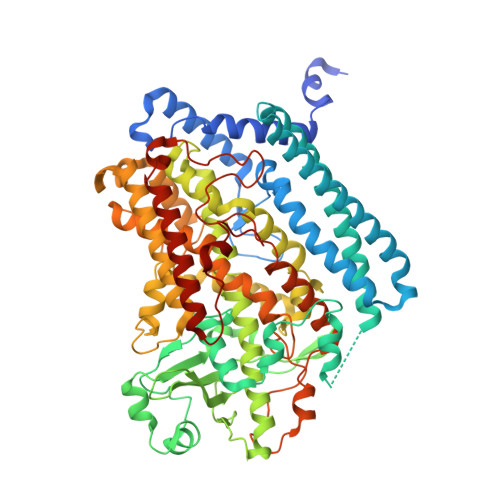
Structural and functional basis of phospholipid oxygenase activity of bacterial lipoxygenase from Pseudomonas aeruginosa.
Banthiya, S., Kalms, J., Galemou Yoga, E., Ivanov, I., Carpena, X., Hamberg, M., Kuhn, H., Scheerer, P.(2016) Biochim Biophys Acta 1861: 1681-1692
- PubMed: 27500637 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbalip.2016.08.002
- Primary Citation Related Structures:
5IR4, 5IR5 - PubMed Abstract:
Pseudomonas aeruginosa expresses a secreted LOX-isoform (PA-LOX, LoxA) capable of oxidizing polyenoic fatty acids to hydroperoxy derivatives. Here we report high-level expression of this enzyme in E. coli and its structural and functional characterization. Recombinant PA-LOX oxygenates polyenoic fatty acids including eicosapentaenoic acid and docosahexaenoic acid to the corresponding (n-6)S-hydroperoxy derivatives. This reaction involves abstraction of the proS-hydrogen from the n-8 bisallylic methylene. PA-LOX lacks major leukotriene synthase activity but converts 5S-HETE and 5S,6R/S-DiHETE to anti-inflammatory and pro-resolving lipoxins. It also exhibits phospholipid oxygenase activity as indicated by the formation of a specific pattern of oxygenation products from different phospholipid subspecies. Multiple mutagenesis studies revealed that PA-LOX does not follow classical concepts explaining the reaction specificity of mammalian LOXs. The crystal structure of PA-LOX was solved with resolutions of up to 1.48Å and its polypeptide chain is folded as single domain. The substrate-binding pocket consists of two fatty acid binding subcavities and lobby. Subcavity-1 contains the catalytic non-heme iron. A phosphatidylethanolamine molecule occupies the substrate-binding pocket and its sn1 fatty acid is located close to the catalytic non-heme iron. His377, His382, His555, Asn559 and the C-terminal Ile685 function as direct iron ligands and a water molecule (hydroxyl) completes the octahedral ligand sphere. Although the biological relevance of PA-LOX is still unknown its functional characteristics (lipoxin synthase activity) implicate this enzyme in a bacterial evasion strategy aimed at downregulating the hosts' immune system.
- Institut für Biochemie, Charité - Universitätsmedizin Berlin, Charitéplatz 1, D-10117 Berlin, Germany.
Organizational Affiliation: