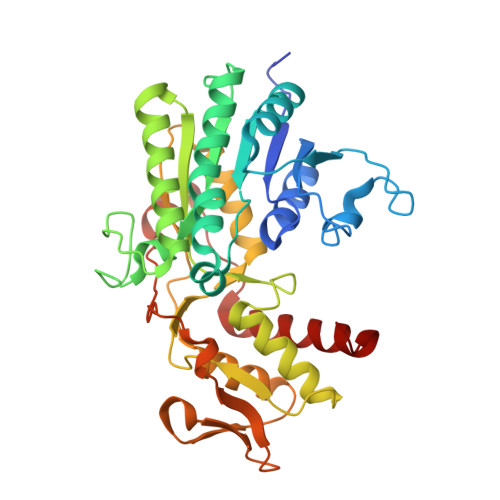
Facile Modulation of Antibody Fucosylation with Small Molecule Fucostatin Inhibitors and Cocrystal Structure with GDP-Mannose 4,6-Dehydratase.
Allen, J.G., Mujacic, M., Frohn, M.J., Pickrell, A.J., Kodama, P., Bagal, D., San Miguel, T., Sickmier, E.A., Osgood, S., Swietlow, A., Li, V., Jordan, J.B., Kim, K.W., Rousseau, A.C., Kim, Y.J., Caille, S., Achmatowicz, M., Thiel, O., Fotsch, C.H., Reddy, P., McCarter, J.D.(2016) ACS Chem Biol 11: 2734-2743
- PubMed: 27434622 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.6b00460
- Primary Citation Related Structures:
5IN4, 5IN5 - PubMed Abstract:
The efficacy of therapeutic antibodies that induce antibody-dependent cellular cytotoxicity can be improved by reduced fucosylation. Consequently, fucosylation is a critical product attribute of monoclonal antibodies produced as protein therapeutics. Small molecule fucosylation inhibitors have also shown promise as potential therapeutics in animal models of tumors, arthritis, and sickle cell disease. Potent small molecule metabolic inhibitors of cellular protein fucosylation, 6,6,6-trifluorofucose per-O-acetate and 6,6,6-trifluorofucose (fucostatin I), were identified that reduces the fucosylation of recombinantly expressed antibodies in cell culture in a concentration-dependent fashion enabling the controlled modulation of protein fucosylation levels. 6,6,6-Trifluorofucose binds at an allosteric site of GDP-mannose 4,6-dehydratase (GMD) as revealed for the first time by the X-ray cocrystal structure of a bound allosteric GMD inhibitor. 6,6,6-Trifluorofucose was found to be incorporated in place of fucose at low levels (<1%) in the glycans of recombinantly expressed antibodies. A fucose-1-phosphonate analog, fucostatin II, was designed that inhibits fucosylation with no incorporation into antibody glycans, allowing the production of afucosylated antibodies in which the incorporation of non-native sugar is completely absent-a key advantage in the production of therapeutic antibodies, especially biosimilar antibodies. Inhibitor structure-activity relationships, identification of cellular and inhibitor metabolites in inhibitor-treated cells, fucose competition studies, and the production of recombinant antibodies with varying levels of fucosylation are described.
- Therapeutic Discovery, Amgen Inc. , One Amgen Center Drive, Thousand Oaks, California 91320, United States.
Organizational Affiliation: