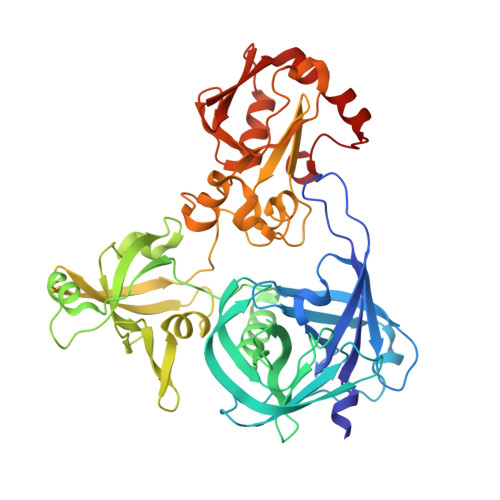
The crystal structure of Deg9 reveals a novel octameric-type HtrA protease
Ouyang, M., Li, X., Zhao, S., Pu, H., Shen, J., Adam, Z., Clausen, T., Zhang, L.(2017) Nat Plants 3: 973-982
- PubMed: 29180814 Search on PubMed
- DOI: https://doi.org/10.1038/s41477-017-0060-2
- Primary Citation Related Structures:
5IL9, 5ILA, 5ILB, 5JYK - PubMed Abstract:
The high temperature requirement A (HtrA) proteases (also termed Deg proteases) play important roles in diverse organisms by regulating protein quality and quantity. One of the 16 Arabidopsis homologs, Deg9, is located in the nucleus where it modulates cytokinin- and light-mediated signalling via degrading the ARABIDOPSIS RESPONSE REGULATOR 4 (ARR4). To uncover the structural features underlying the proteolytic activity of Deg9, we determined its crystal structure. Unlike the well-established trimeric building block of HtrAs, Deg9 displays a novel octameric structure consisting of two tetrameric rings that have distinct conformations. Based on the structural architecture, we generated several mutant variants of Deg9, determined their structure and tested their proteolytic activity towards ARR4. The results of the structural and biochemical analyses allowed us to propose a model for a novel mechanism of substrate recognition and activity regulation of Deg9. In this model, protease activation of one tetramer is mediated by en-bloc reorientation of the protease domains to open an entrance for the substrate in the opposite (inactive) tetramer. This study provides the structural basis for understanding how the levels of nuclear signal components are regulated by a plant protease.
- Photosynthesis Research Center, Key Laboratory of Photobiology, Institute of Botany, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: