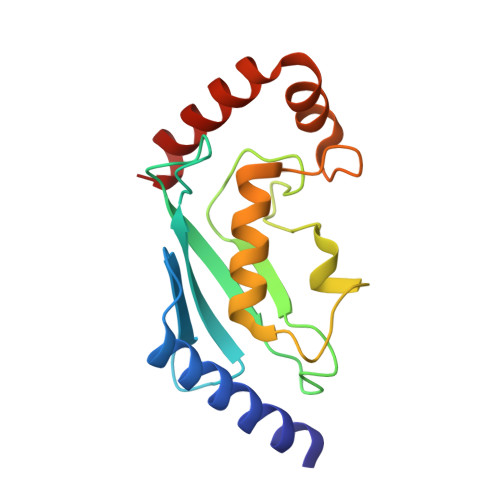
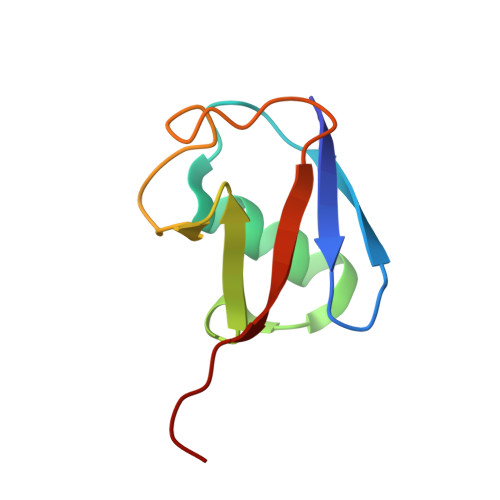
A cascading activity-based probe sequentially targets E1-E2-E3 ubiquitin enzymes.
Mulder, M.P., Witting, K., Berlin, I., Pruneda, J.N., Wu, K.P., Chang, J.G., Merkx, R., Bialas, J., Groettrup, M., Vertegaal, A.C., Schulman, B.A., Komander, D., Neefjes, J., El Oualid, F., Ovaa, H.(2016) Nat Chem Biol 12: 523-530
- PubMed: 27182664 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.2084
- Primary Citation Related Structures:
5IFR - PubMed Abstract:
Post-translational modifications of proteins with ubiquitin (Ub) and ubiquitin-like modifiers (Ubls), orchestrated by a cascade of specialized E1, E2 and E3 enzymes, control a wide range of cellular processes. To monitor catalysis along these complex reaction pathways, we developed a cascading activity-based probe, UbDha. Similarly to the native Ub, upon ATP-dependent activation by the E1, UbDha can travel downstream to the E2 (and subsequently E3) enzymes through sequential trans-thioesterifications. Unlike the native Ub, at each step along the cascade, UbDha has the option to react irreversibly with active site cysteine residues of target enzymes, thus enabling their detection. We show that our cascading probe 'hops' and 'traps' catalytically active Ub-modifying enzymes (but not their substrates) by a mechanism diversifiable to Ubls. Our founder methodology, amenable to structural studies, proteome-wide profiling and monitoring of enzymatic activity in living cells, presents novel and versatile tools to interrogate Ub and Ubl cascades.
- Division of Cell Biology, Netherlands Cancer Institute, Amsterdam, the Netherlands.
Organizational Affiliation: