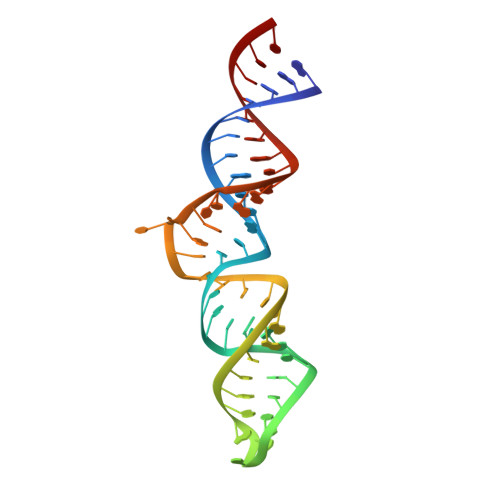
Solution structure of the 5'-terminal hairpin of the 7SK small nuclear RNA.
Bourbigot, S., Dock-Bregeon, A.C., Eberling, P., Coutant, J., Kieffer, B., Lebars, I.(2016) RNA 22: 1844-1858
- PubMed: 27852926
- DOI: https://doi.org/10.1261/rna.056523.116
- Primary Citation of Related Structures:
5IEM - PubMed Abstract:
The small nuclear 7SK RNA regulates RNA polymerase II (RNA Pol II) transcription, by sequestering and inhibiting the positive transcription elongation factor b (P-TEFb). P-TEFb is stored in the 7SK ribonucleoprotein (RNP) that contains the three nuclear proteins Hexim1, LaRP7, and MePCE. P-TEFb interacts with the protein Hexim1 and the 7SK RNA. Once P-TEFb is released from the 7SK RNP, it activates transcription by phosphorylating the C-terminal domain of RNA Pol II. P-TEFb also plays a crucial role in the replication of the human immunodeficiency virus HIV-1, through its recruitment by the viral transactivator Tat. Previous work demonstrated that the protein Tat promotes the release of P-TEFb from the 7SK RNP through direct binding to the 7SK RNA. Hexim1 and Tat proteins both comprise conserved and similar arginine-rich motifs that were identified to bind the 7SK RNA at a repeated GAUC site located at the top of the 5'-terminal hairpin (HPI). Here, we report the solution structure of this region as determined by nuclear magnetic resonance, to identify HPI structural features recognized by Hexim1 and Tat. The HPI solution structure displays an elongated shape featuring four helical segments interrupted by one internal loop and three bulges with distinct folds. In particular, the repeated GAUC motif adopts a pre-organized geometry. Our results suggest that the binding of Hexim1 and Tat to the 7SK RNA could originate from a conformational selection of this motif, highlighting how RNA local structure could lead to an adaptive recognition of their partners.
- Department of Integrated Structural Biology, Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), CNRS UMR 7104, INSERM U964, Université de Strasbourg, 67404 Illkirch Cedex, France.
Organizational Affiliation: