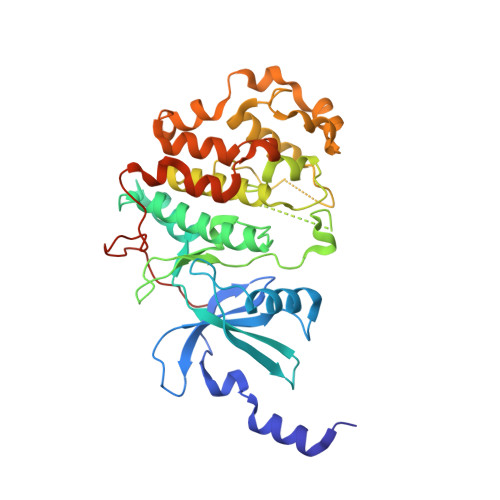
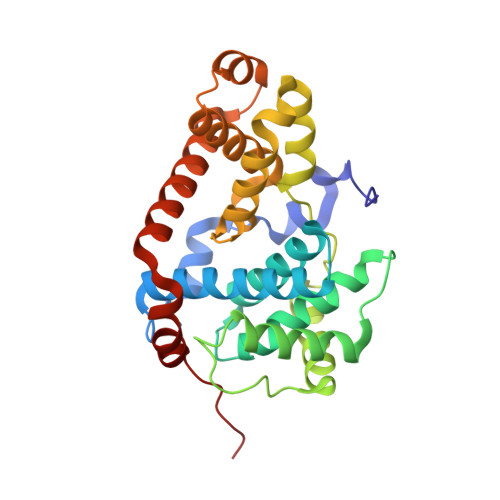
Structure-Based Optimization of Potent, Selective, and Orally Bioavailable CDK8 Inhibitors Discovered by High-Throughput Screening.
Czodrowski, P., Mallinger, A., Wienke, D., Esdar, C., Poschke, O., Busch, M., Rohdich, F., Eccles, S.A., Ortiz-Ruiz, M.J., Schneider, R., Raynaud, F.I., Clarke, P.A., Musil, D., Schwarz, D., Dale, T., Urbahns, K., Blagg, J., Schiemann, K.(2016) J Med Chem 59: 9337-9349
- PubMed: 27490956 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00597
- Primary Citation Related Structures:
5ICP, 5IDN, 5IDP - PubMed Abstract:
The mediator complex-associated cyclin dependent kinase CDK8 regulates β-catenin-dependent transcription following activation of WNT signaling. Multiple lines of evidence suggest CDK8 may act as an oncogene in the development of colorectal cancer. Here we describe the successful optimization of an imidazo-thiadiazole series of CDK8 inhibitors that was identified in a high-throughput screening campaign and further progressed by structure-based design. In several optimization cycles, we improved the microsomal stability, potency, and kinase selectivity. The initial imidazo-thiadiazole scaffold was replaced by a 3-methyl-1H-pyrazolo[3,4-b]-pyridine which resulted in compound 25 (MSC2530818) that displayed excellent kinase selectivity, biochemical and cellular potency, microsomal stability, and is orally bioavailable. Furthermore, we demonstrated modulation of phospho-STAT1, a pharmacodynamic biomarker of CDK8 activity, and tumor growth inhibition in an APC mutant SW620 human colorectal carcinoma xenograft model after oral administration. Compound 25 demonstrated suitable potency and selectivity to progress into preclinical in vivo efficacy and safety studies.
- Merck KGaA , Frankfurter Strasse 250, Darmstadt, 64293, Germany.
Organizational Affiliation: