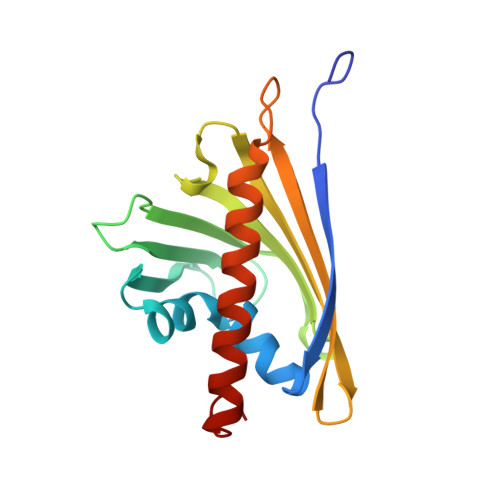
Crystal Structure of Hyp-1, a Hypericum perforatum PR-10 Protein, in Complex with Melatonin.
Sliwiak, J., Dauter, Z., Jaskolski, M.(2016) Front Plant Sci 7: 668-668
- PubMed: 27242869 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fpls.2016.00668
- Primary Citation Related Structures:
5I8F - PubMed Abstract:
Hyp-1, a PR-10-fold protein from Hypericum perforatum, was crystallized in complex with melatonin (MEL). The structure confirms the conserved protein fold and the presence of three unusual ligand binding sites, two of which are internal chambers (1,2), while the third one (3) is formed as an invagination of the protein surface. The MEL ligand in site 1 is well defined while that in site 3 seems to be rotating between the side chains of Lys33 and Tyr150 that act as a molecular vise. The patch of electron density in site 2 does not allow unambiguous modeling of a melatonin molecule but suggests a possible presence of its degradation product. This pattern of ligand occupation is reproducible in repeated crystallization/structure determination experiments. Although the binding of melatonin by Hyp-1 does not appear to be very strong (for example, MEL cannot displace the artificial fluorescence probe ANS), it is strong enough to suggest a physiological role of this interaction. For example, trans-zeatin, which is a common ligand of PR-10 proteins, does not overcompete melatonin for binding to Hyp-1 as it does not affect the crystallization process of the Hyp-1/MEL complex, and among a number of potential natural mediators tested, melatonin was the only one to form a crystalline complex with Hyp-1 with the use of standard crystallization screens. Hyp-1 is the second protein in the Protein Data Bank for which melatonin binding has been demonstrated crystallographically, the first one being human quinone reductase.
- Center for Biocrystallographic Research, Institute of Bioorganic Chemistry, Polish Academy of Sciences Poznan, Poland.
Organizational Affiliation: