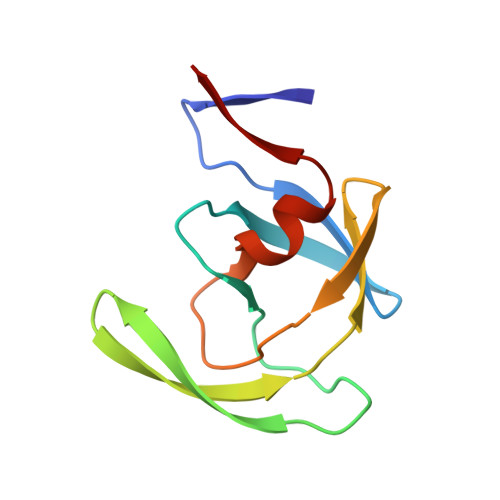
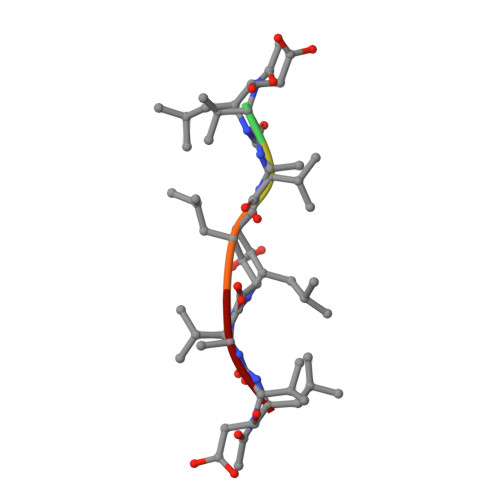
Crystallographic analysis of a complex between human immunodeficiency virus type 1 protease and acetyl-pepstatin at 2.0-A resolution.
Fitzgerald, P.M., McKeever, B.M., VanMiddlesworth, J.F., Springer, J.P., Heimbach, J.C., Leu, C.T., Herber, W.K., Dixon, R.A., Darke, P.L.(1990) J Biological Chem 265: 14209-14219
- PubMed: 2201682 Search on PubMed
- Primary Citation Related Structures:
5HVP - PubMed Abstract:
The mode of binding of acetyl-pepstatin to the protease from the human immunodeficiency virus type 1 (HIV-1) has been determined by x-ray diffraction analysis. Crystals of an acetyl-pepstatin-HIV-1 protease complex were obtained in space group P2(1)2(1)2 (unit cell dimensions a = 58.39 A, b = 86.70 A, c = 46.27 A) by precipitation with sodium chloride. The structure was phased by molecular replacement methods, and a model for the structure was refined using diffraction data to 2.0 A resolution (R = 0.176 for 12901 reflections with I greater than sigma (I); deviation of bond distances from ideal values = 0.018 A; 172 solvent molecules included). The structure of the protein in the complex has been compared with the structure of the enzyme without the ligand. A core of 44 amino acids in each monomer, including residues in the active site and residues at the dimer interface, remains unchanged on binding of the inhibitor (root mean square deviation of alpha carbon positions = 0.39 A). The remaining 55 residues in each monomer undergo substantial rearrangement, with the most dramatic changes occurring at residues 44-57 (these residues comprise the so-called flaps of the enzyme). The flaps interact with one another and with the inhibitor so as to largely preserve the 2-fold symmetry of the protein. The inhibitor is bound in two approximately symmetric orientations. In both orientations the peptidyl backbone of the inhibitor is extended; a network of hydrogen bonds is formed between the inhibitor and the main body of the protein as well as between the inhibitor and the flaps. Hydrophobic side chains of residues in the body of the protein form partial binding sites for the side chains of the inhibitor; hydrophobic side chains of residues in the flaps complete these binding sites.
- Department of Biophysical Chemistry, Merck Sharp and Dohme Research Laboratories, Rahway, New Jersey 07065.
Organizational Affiliation: