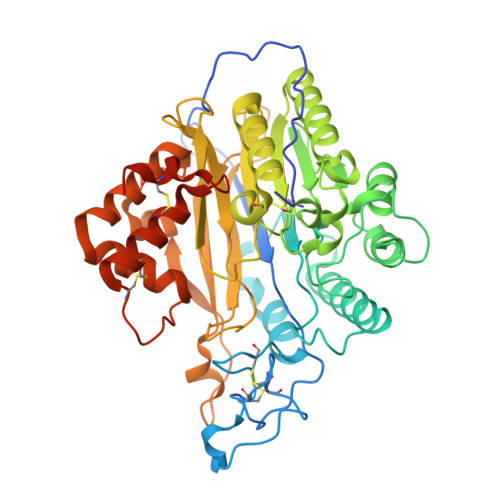
Crystal structure of mammalian acid sphingomyelinase.
Gorelik, A., Illes, K., Heinz, L.X., Superti-Furga, G., Nagar, B.(2016) Nat Commun 7: 12196-12196
- PubMed: 27435900
- DOI: https://doi.org/10.1038/ncomms12196
- Primary Citation of Related Structures:
5FI9, 5FIB, 5FIC, 5HQN - PubMed Abstract:
Acid sphingomyelinase (ASMase, ASM, SMPD1) converts sphingomyelin into ceramide, modulating membrane properties and signal transduction. Inactivating mutations in ASMase cause Niemann-Pick disease, and its inhibition is also beneficial in models of depression and cancer. To gain a better understanding of this critical therapeutic target, we determined crystal structures of mammalian ASMase in various conformations. The catalytic domain adopts a calcineurin-like fold with two zinc ions and a hydrophobic track leading to the active site. Strikingly, the membrane interacting saposin domain assumes either a closed globular conformation independent from the catalytic domain, or an open conformation, which establishes an interface with the catalytic domain essential for activity. Structural mapping of Niemann-Pick mutations reveals that most of them likely destabilize the protein's fold. This study sheds light on the molecular mechanism of ASMase function, and provides a platform for the rational development of ASMase inhibitors and therapeutic use of recombinant ASMase.
- Department of Biochemistry, McGill University, Montreal, Quebec, Canada H3G 0B1.
Organizational Affiliation: