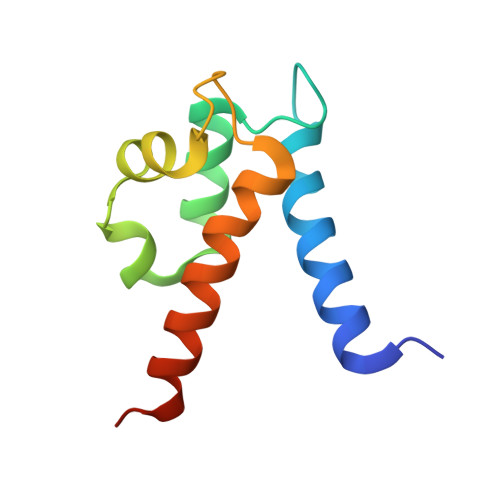
Crystal structure of human S100A8 in complex with zinc and calcium.
Lin, H., Andersen, G.R., Yatime, L.(2016) BMC Struct Biol 16: 8-8
- PubMed: 27251136 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s12900-016-0058-4
- Primary Citation Related Structures:
5HLO, 5HLV - PubMed Abstract:
S100 proteins are a large family of calcium binding proteins present only in vertebrates. They function intra- and extracellularly both as regulators of homeostatic processes and as potent effectors during inflammation. Among these, S100A8 and S100A9 are two major constituents of neutrophils that can assemble into homodimers, heterodimers and higher oligomeric species, including fibrillary structures found in the ageing prostate. Each of these forms assumes specific functions and their formation is dependent on divalent cations, notably calcium and zinc. In particular, zinc appears as a major regulator of S100 protein function in a disease context. Despite this central role, no structural information on how zinc bind to S100A8/S100A9 and regulates their quaternary structure is yet available. Here we report two crystallographic structures of calcium and zinc-loaded human S100A8. S100A8 binds two zinc ions per homodimer, through two symmetrical, all-His tetracoordination sites, revealing a classical His-Zn binding mode for the protein. Furthermore, the presence of a (Zn)2-cacodylate complex in our second crystal form induces ligand swapping within the canonical His4 zinc binding motif, thereby creating two new Zn-sites, one of which involves residues from symmetry-related molecules. Finally, we describe the calcium-induced S100A8 tetramer and reveal how zinc stabilizes this tetramer by tightening the dimer-dimer interface. Our structures of Zn(2+)/Ca(2+)-bound hS100A8 demonstrate that S100A8 is a genuine His-Zn S100 protein. Furthermore, they show how zinc stabilizes S100A8 tetramerization and potentially mediates the formation of novel interdimer interactions. We propose that these zinc-mediated interactions may serve as a basis for the generation of larger oligomers in vivo.
- Department of Molecular Biology and Genetics, Aarhus University, Gustav Wieds Vej 10C, DK-8000, Aarhus, Denmark.
Organizational Affiliation: