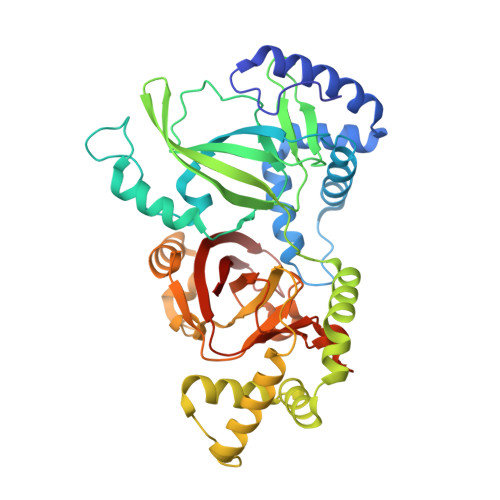
Crystal structure and structure-based mutagenesis of actin-specific ADP-ribosylating toxin CPILE-a as novel enterotoxin
Toniti, W., Yoshida, T., Tsurumura, T., Irikura, D., Monma, C., Kamata, Y., Tsuge, H.(2017) PLoS One 12: e0171278-e0171278
- PubMed: 28199340
- DOI: https://doi.org/10.1371/journal.pone.0171278
- Primary Citation Related Structures:
5GTT, 5WTZ, 5WU0 - PubMed Abstract:
Unusual outbreaks of food poisoning in Japan were reported in which Clostridium perfringens was strongly suspected to be the cause based on epidemiological information and fingerprinting of isolates. The isolated strains lack the typical C. perfringens enterotoxin (CPE) but secrete a new enterotoxin consisting of two components: C. perfringens iota-like enterotoxin-a (CPILE-a), which acts as an enzymatic ADP-ribosyltransferase, and CPILE-b, a membrane binding component. Here we present the crystal structures of apo-CPILE-a, NAD+-CPILE-a and NADH-CPILE-a. Though CPILE-a structure has high similarity with known iota toxin-a (Ia) with NAD+, it possesses two extra-long protruding loops from G262-S269 and E402-K408 that are distinct from Ia. Based on the Ia-actin complex structure, we focused on actin-binding interface regions (I-V) including two protruding loops (PT) and examined how mutations in these regions affect the ADP-ribosylation activity of CPILE-a. Though some site-directed mutagenesis studies have already been conducted on the actin binding site of Ia, in the present study, mutagenesis studies were conducted against both α- and β/γ-actin in CPILE-a and Ia. Interestingly, CPILE-a ADP-ribosylates both α- and β/γ-actin, but its sensitivity towards β/γ-actin is 36% compared with α-actin. Our results contrast to that only C2-I ADP-ribosylates β/γ-actin. We also showed that PT-I and two convex-concave interactions in CPILE-a are important for actin binding. The current study is the first detailed analysis of site-directed mutagenesis in the actin binding region of Ia and CPILE-a against both α- and β/γ-actin.
- Department of Bioresource and Environmental Sciences, Faculty of Life Sciences, Kyoto Sangyo University, Kyoto, Japan.
Organizational Affiliation: