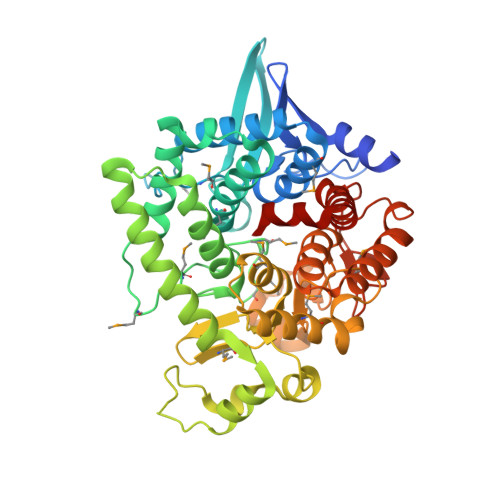
Structural Analysis of the Catalytic Mechanism and Substrate Specificity of Anabaena Alkaline Invertase InvA Reveals a Novel Glucosidase
Xie, J., Cai, K., Hu, H.X., Jiang, Y.L., Yang, F., Hu, P.F., Cao, D.D., Li, W.F., Chen, Y., Zhou, C.Z.(2016) J Biological Chem 291: 25667-25677
- PubMed: 27777307
- DOI: https://doi.org/10.1074/jbc.M116.759290
- Primary Citation Related Structures:
5GOO, 5GOP, 5GOQ, 5GOR - PubMed Abstract:
Invertases catalyze the hydrolysis of sucrose to glucose and fructose, thereby playing a key role in primary metabolism and plant development. According to the optimum pH, invertases are classified into acid invertases (Ac-Invs) and alkaline/neutral invertases (A/N-Invs), which share no sequence homology. Compared with Ac-Invs that have been extensively studied, the structure and catalytic mechanism of A/N-Invs remain unknown. Here we report the crystal structures of Anabaena alkaline invertase InvA, which was proposed to be the ancestor of modern plant A/N-Invs. These structures are the first in the GH100 family. InvA exists as a hexamer in both crystal and solution. Each subunit consists of an (α/α) 6 barrel core structure in addition to an insertion of three helices. A couple of structures in complex with the substrate or products enabled us to assign the subsites -1 and +1 specifically binding glucose and fructose, respectively. Structural comparison combined with enzymatic assays indicated that Asp-188 and Glu-414 are putative catalytic residues. Further analysis of the substrate binding pocket demonstrated that InvA possesses a stringent substrate specificity toward the α1,2-glycosidic bond of sucrose. Together, we suggest that InvA and homologs represent a novel family of glucosidases.
- From the Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei Anhui 230027, China.
Organizational Affiliation: