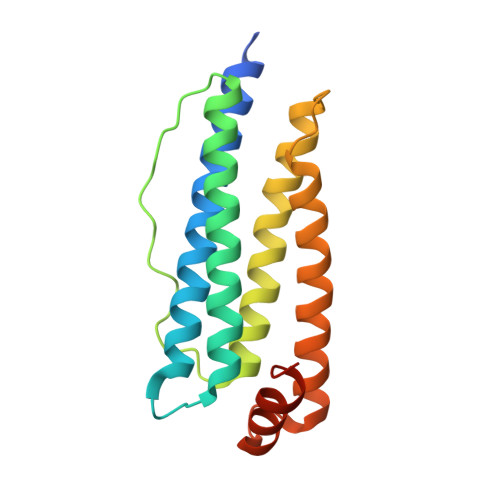
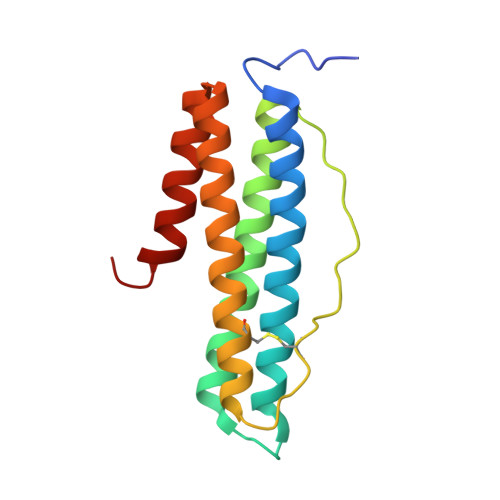
"Silent" Amino Acid Residues at Key Subunit Interfaces Regulate the Geometry of Protein Nanocages
Zhang, S., Zang, J., Zhang, X., Chen, H., Mikami, B., Zhao, G.(2016) ACS Nano 10: 10382-10388
- PubMed: 27934076 Search on PubMed
- DOI: https://doi.org/10.1021/acsnano.6b06235
- Primary Citation Related Structures:
5GN8 - PubMed Abstract:
Rendering the geometry of protein-based assemblies controllable remains challenging. Protein shell-like nanocages represent particularly interesting targets for designed assembly. Here, we introduce an engineering strategy-key subunit interface redesign (KSIR)-that alters a natural subunit-subunit interface by selective deletion of a small number of "silent" amino acid residues (no participation in interfacial interactions) into one that triggers the generation of a non-native protein cage. We have applied KSIR to construct a non-native 48-mer nanocage from its native 24-mer recombinant human H-chain ferritin (rHuHF). This protein is a heteropolymer composed of equal numbers of two different subunits which are derived from one polypeptide. This strategy has allowed the study of conversion between protein nanocages with different geometries by re-engineering key subunit interfaces and the demonstration of the important role of the above-mentioned specific residues in providing geometric specificity for protein assembly.
- Beijing Advanced Innovation Center for Food Nutrition and Human Health, College of Food Science and Nutritional Engineering, China Agricultural University , Key Laboratory of Functional Dairy, Ministry of Education, Beijing 100083, China.
Organizational Affiliation: