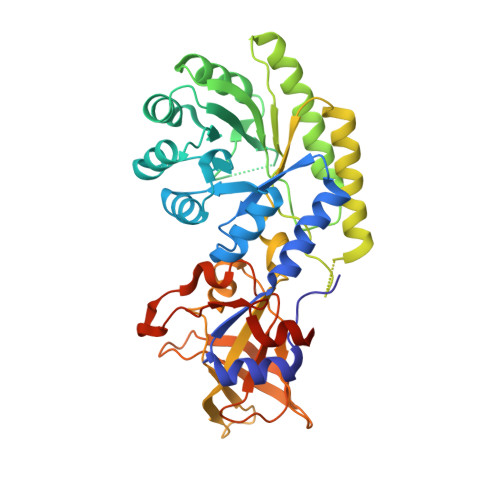
Crystal Structure and Pyridoxal 5-Phosphate Binding Property of Lysine Decarboxylase from Selenomonas ruminantium
Sagong, H.-Y., Son, H.F., Kim, S., Kim, Y.-H., Kim, I.-K., Kim, K.-J.(2016) PLoS One 11: e0166667-e0166667
- PubMed: 27861532
- DOI: https://doi.org/10.1371/journal.pone.0166667
- Primary Citation of Related Structures:
5GJM, 5GJN, 5GJP - PubMed Abstract:
Lysine decarboxylase (LDC) is a crucial enzyme for acid stress resistance and is also utilized for the biosynthesis of cadaverine, a promising building block for bio-based polyamides. We determined the crystal structure of LDC from Selenomonas ruminantium (SrLDC). SrLDC functions as a dimer and each monomer consists of two distinct domains; a PLP-binding barrel domain and a sheet domain. We also determined the structure of SrLDC in complex with PLP and cadaverine and elucidated the binding mode of cofactor and substrate. Interestingly, compared with the apo-form of SrLDC, the SrLDC in complex with PLP and cadaverine showed a remarkable structural change at the PLP binding site. The PLP binding site of SrLDC contains the highly flexible loops with high b-factors and showed an open-closed conformational change upon the binding of PLP. In fact, SrLDC showed no LDC activity without PLP supplement, and we suggest that highly flexible PLP binding site results in low PLP affinity of SrLDC. In addition, other structurally homologous enzymes also contain the flexible PLP binding site, which indicates that high flexibility at the PLP binding site and low PLP affinity seems to be a common feature of these enzyme family.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daegu, 702-701, Republic of Korea.
Organizational Affiliation: