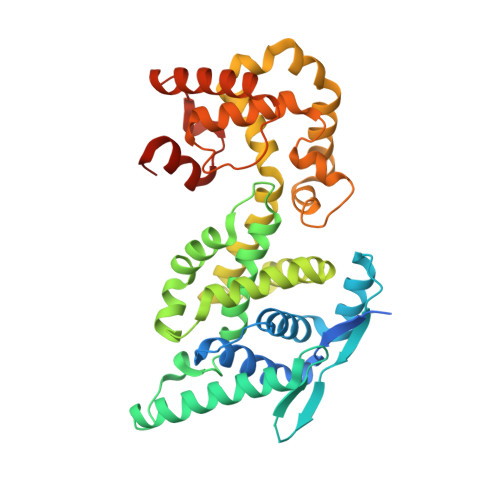
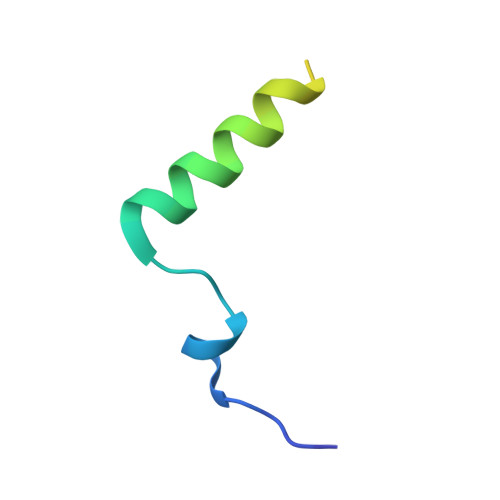
Nucleocapsid assembly in pneumoviruses is regulated by conformational switching of the N protein.
Renner, M., Bertinelli, M., Leyrat, C., Paesen, G.C., Saraiva de Oliveira, L.F., Huiskonen, J.T., Grimes, J.M.(2016) Elife 5: e12627-e12627
- PubMed: 26880565 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.12627
- Primary Citation Related Structures:
5FVC, 5FVD - PubMed Abstract:
Non-segmented, (-)RNA viruses cause serious human diseases. Human metapneumovirus (HMPV), an emerging pathogen of this order of viruses (Mononegavirales) is one of the main causes of respiratory tract illness in children. To help elucidate the assembly mechanism of the nucleocapsid (the viral RNA genome packaged by the nucleoprotein N) we present crystallographic structures of HMPV N in its assembled RNA-bound state and in a monomeric state, bound to the polymerase cofactor P. Our structures reveal molecular details of how P inhibits the self-assembly of N and how N transitions between the RNA-free and RNA-bound conformational state. Notably, we observe a role for the C-terminal extension of N in directly preventing premature uptake of RNA by folding into the RNA-binding cleft. Our structures suggest a common mechanism of how the growth of the nucleocapsid is orchestrated, and highlight an interaction site representing an important target for antivirals.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, United Kingdom.
Organizational Affiliation: