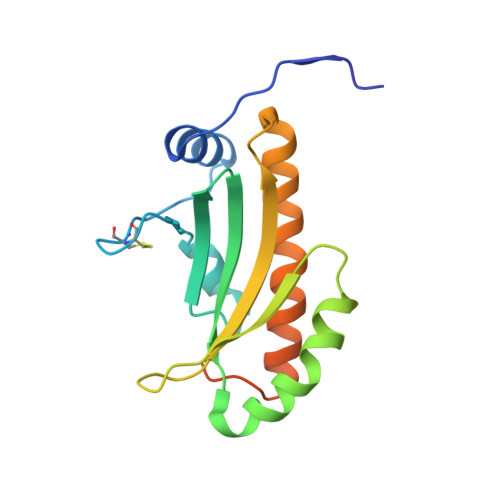
The crystal structure of Erwinia amylovora AmyR, a member of the YbjN protein family, shows similarity to type III secretion chaperones but suggests different cellular functions.
Bartho, J.D., Bellini, D., Wuerges, J., Demitri, N., Toccafondi, M., Schmitt, A.O., Zhao, Y., Walsh, M.A., Benini, S.(2017) PLoS One 12: e0176049-e0176049
- PubMed: 28426806
- DOI: https://doi.org/10.1371/journal.pone.0176049
- Primary Citation Related Structures:
5FR7, 5FRK - PubMed Abstract:
AmyR is a stress and virulence associated protein from the plant pathogenic Enterobacteriaceae species Erwinia amylovora, and is a functionally conserved ortholog of YbjN from Escherichia coli. The crystal structure of E. amylovora AmyR reveals a class I type III secretion chaperone-like fold, despite the lack of sequence similarity between these two classes of protein and lacking any evidence of a secretion-associated role. The results indicate that AmyR, and YbjN proteins in general, function through protein-protein interactions without any enzymatic action. The YbjN proteins of Enterobacteriaceae show remarkably low sequence similarity with other members of the YbjN protein family in Eubacteria, yet a high level of structural conservation is observed. Across the YbjN protein family sequence conservation is limited to residues stabilising the protein core and dimerization interface, while interacting regions are only conserved between closely related species. This study presents the first structure of a YbjN protein from Enterobacteriaceae, the most highly divergent and well-studied subgroup of YbjN proteins, and an in-depth sequence and structural analysis of this important but poorly understood protein family.
- Bioorganic Chemistry and Bio-Crystallography laboratory (B2Cl), Faculty of Science and Technology, Free University of Bolzano, Piazza Università 5, Bolzano, Italy.
Organizational Affiliation: