Structural and Functional Investigations of the Effector Protein LpiR1 from Legionella pneumophila.
Beyrakhova, K.A., van Straaten, K., Li, L., Boniecki, M.T., Anderson, D.H., Cygler, M.(2016) J Biological Chem 291: 15767-15777
- PubMed: 27226543 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.708701
- Primary Citation Related Structures:
5FIA, 5JG4 - PubMed Abstract:
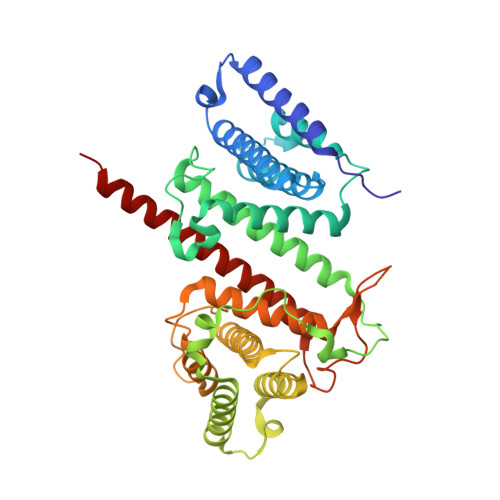
Legionella pneumophila is a causative agent of a severe pneumonia, known as Legionnaires' disease. Legionella pathogenicity is mediated by specific virulence factors, called bacterial effectors, which are injected into the invaded host cell by the bacterial type IV secretion system. Bacterial effectors are involved in complex interactions with the components of the host cell immune and signaling pathways, which eventually lead to bacterial survival and replication inside the mammalian cell. Structural and functional studies of bacterial effectors are, therefore, crucial for elucidating the mechanisms of Legionella virulence. Here we describe the crystal structure of the LpiR1 (Lpg0634) effector protein and investigate the effects of its overexpression in mammalian cells. LpiR1 is an α-helical protein that consists of two similar domains aligned in an antiparallel fashion. The hydrophilic cleft between the domains might serve as a binding site for a potential host cell interaction partner. LpiR1 binds the phosphate group at a conserved site and is stabilized by Mn(2+), Ca(2+), or Mg(2+) ions. When overexpressed in mammalian cells, a GFP-LpiR1 fusion protein is localized in the cytoplasm. Intracellular signaling antibody array analysis revealed small changes in the phosphorylation state of several components of the Akt signaling pathway in HEK293T cells overexpressing LpiR1.
- From the Department of Biochemistry and.
Organizational Affiliation: